Pollen contamination and mating patterns in a Prosopis alba clonal orchard: impact on seed orchards establishment
iForest - Biogeosciences and Forestry, Volume 12, Issue 3, Pages 330-337 (2019)
doi: https://doi.org/10.3832/ifor2936-012
Published: Jun 14, 2019 - Copyright © 2019 SISEF
Research Articles
Abstract
Prosopis alba (Leguminosae) is an important species from ecologic and economical points of view in arid and semi-arid regions of Argentina. In several open-pollinated species, pollen contamination from off-orchard parents and selfing have been proven to reduce orchard seed quality. In 2002, the first clonal orchard of Prosopis alba was established in Fernández (Santiago del Estero, Argentina) with 12 trees phenotypically selected from a progeny trial, based on height, pod production per year and pod sweetness. The aim of this study was to evaluate the mating patterns and pollen contamination rate in the orchard using ten SSR markers and paternity analysis. All the clones together with the progeny of a single clone (open-pollinated seeds) were genotyped. Data was processed by two different methods based on likelihood and Bayesian approaches, respectively. A high consistency (89%) of results was observed between the two methods, and pollen contamination rate was estimated between 27% and 37%. The minimum number of different pollen donors per mother plant varied from three to five and selfing occurrence was low (<1.6%). Based on the estimated status number (Ns = 4.4), the expected coancestry in the seed crop is equal to a Mendelian population with an effective size of 4-5 individuals. Genetic analyses are encouraged during the establishment and monitoring of trials in forest breeding and management programmes. It is strongly recommended to establish seed orchards in isolated areas and to guarantee equal representation of parental genotypes in the orchards.
Keywords
Cervus, MasterBayes, Microsatellites, Paternity Analysis, Mesquite, Prosopis alba, Seed Orchard
Introduction
Prosopis alba is an important ecological and economical species distributed in arid and semiarid areas of South America, belonging to genus Prosopis (Leguminosae -[11]). It is a native forest tree widely found in Argentina, Uruguay, Paraguay and from south of Bolivia to Peru. Nowadays, overexploitation and land conversion into crop plantations in the biogeographical Chaco region have led to important loss of its forest habitats. The large genetic differentiation among P. alba populations and its distribution in a variety of different environments may be exploited in forest tree breeding programs of this species. Previous studies indicated that Prosopis species are mainly outcrossers ([3]) with endozoic seed dispersion mediated by small and medium mammals ([39]), while pollen is mainly dispersed by insects ([20]). In a natural population of P. alba the pollen was estimated to disperse from 5 to 30 m around each tree ([3]). Consistently, studies on the spatial genetic structure indicated that the dispersal distance in P. alba ranges from 66 to 250 m ([7]).
Seed orchards are efficient tools to produce seeds easily, abundantly and economically from trees with desirable genetic properties ([13]). They can be constituted from seeds (sexual orchards) or from vegetative propagules from selected individuals based on phenotypic traits (clonal orchard - [12]). Their efficiency depends on the effective size of the orchard and the ability to exclude non-selected individuals ([46]). Ideally, seed orchards are designed to avoid selfing and outer pollen contamination, assuming that all crosses are equally available and compatible ([9]). Pollen contamination by non-selected trees from outside of the orchards and selfing may have similar negative effects on the progeny by reducing the genetic gain ([32]). Pollen contamination level depends on the amount of pollen produced by the selected trees and their geographic isolation from external non-desired sources.
An accurate estimation of seed orchard reproductive patterns are necessary. Parental analyses have become an important method for providing a detail description of genetic diversity and pollination patterns inside the seed orchards mainly in anemophilous tree species ([23], [43], [19], [8], [14]). However, for insect pollinated trees only few studies are available ([42], [48]). Simple sequence repeats (SSRs) are the most used markers for parental analysis. Both likelihood and Bayesian approaches ([25], [42]) showed to be useful for parental assignments in forest tree species.
Currently, P. alba breeding programmes imply the establishment of several provenance trials in different sites of Argentina together with a few clonal orchards ([18], [31], [16], [17]). Although origin certified seeds are available from INASE (Instituto Nacional de Semillas - National Institute of Seeds), seeds from provenance trials have not quality certification yet. In 1990, a progeny trial was established in San Carlos, Santiago del Estero (Argentina), in a randomized complete block design with 57 families, 7 replication and 4 trees per replication. This progeny trial was established from seeds representing wild open-pollinated families from mother plants with good phenotypic characteristics, considering vigor, crown size, unbranched condition, long and straight trunk, and healthy aspect ([18]). The original stand included samples from eight unrelated provenances representing different areas and P. alba morphotypes. The Santiagueño type was represented by the populations of Rio Dulce Irrigation Zone (RDU), Sumampa (SUM), Pinto (PIN) and Añatuya (ANA), the Chaco Norte by Castelli (CAS) and Ibarreta (IBA) and the Chaco Sur type by Quimili (QUI) and Gato Colorado (GCO - [6]). In 1999, 12 trees were selected from this trial, based on height, pod production per year and pod sweetness. The selected trees belong to different families and derived from the 4 provenances RDU and PIN from Santiago del Estero, CAS from Chaco, and IBA from Formosa (represented respectively by 4, 1, 4, and 3 trees). From these trees a clonal orchard was established in 2002 at the Estación Experimental Fernández (Convenio Universidad Católica de Santiago del Estero, Santiago del Estero, Argentina - [18]).
The purpose of the present study was to estimate pollen contamination rate and pollination patterns in this first-generation clonal orchard of P. alba using SSR markers, in order to provide significant information for developing clonal seed orchard management strategies and evaluating the impact of pollen contamination on the expected gain.
Materials and methods
Seed orchard and plant material
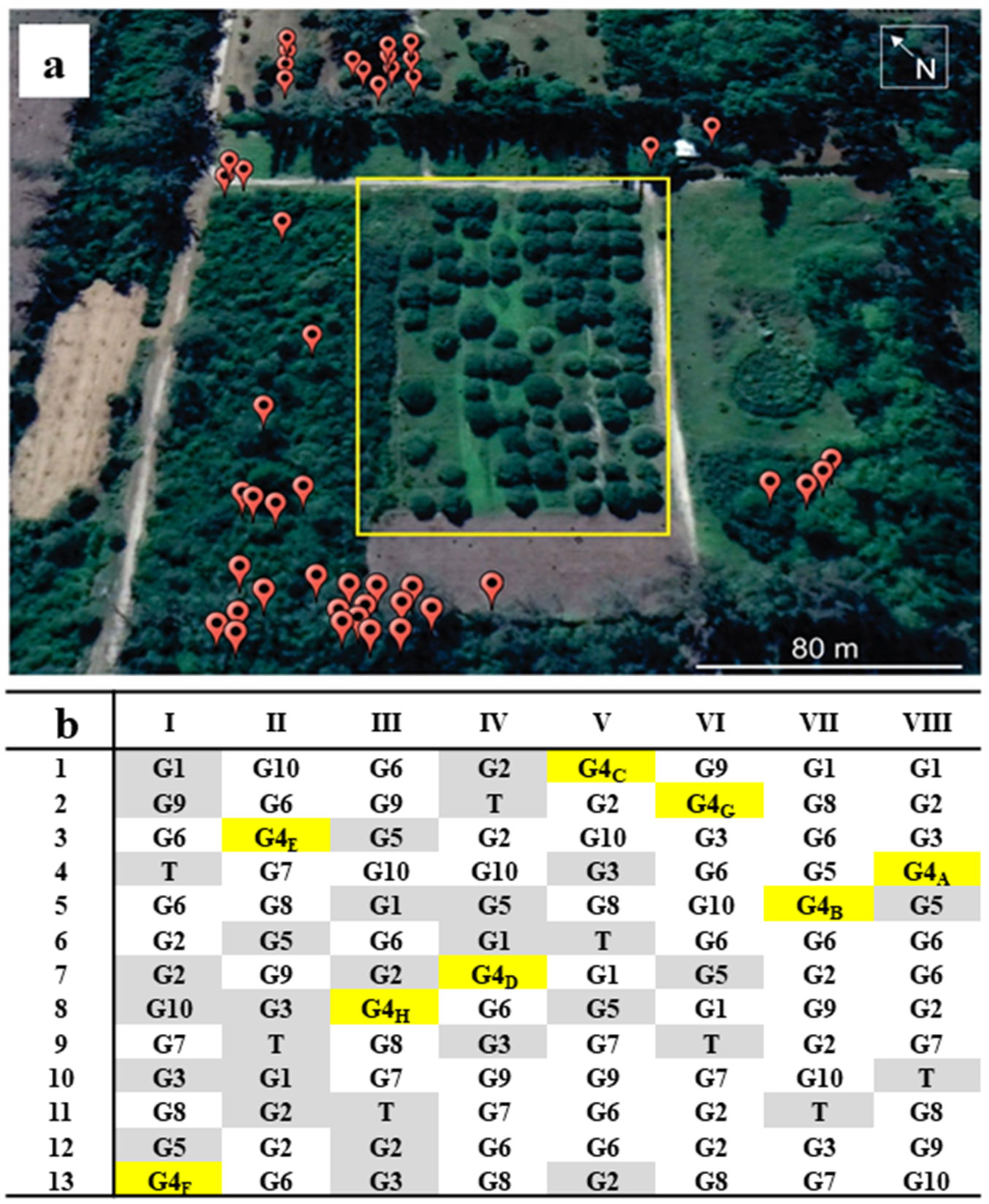
A P. alba progeny trial was established in 1990, 10 km East of Santiago del Estero City, Argentina (27 ° 45′ S; 64 ° 12′ W). The experimental design was a randomized complete block design with 57 families, seven replications and four trees per replication. From them, 12 trees were selected based on height, pod production and pod sweetness and cloned ([18]). Shoots from these clones were multiplied by rooting of cuttings and implanted in 2002 in a new clonal orchard situated at the Estación Experimental Fernández (Convenio Universidad Católica de Santiago del Estero, Santiago del Estero, Argentina - 27° 56′ S, 65° 52.5′ W) using 10 × 10 m spacing between trees. The orchard was constituted by 8 completely randomized blocks, including ramets of the 12 selected clones and one reference tree (non selected) in each block, making a total of 104 plantlets (Fig. 1). However, the design became unbalanced because at the implantation time the number of cuttings that were successfully rooted and developed varied among the clones reducing the effective number of installed trees (M. Ewens, personal communication). Naturally regenerated trees in the clonal orchard site were not allowed. In the surrounding area, 47 P. alba plants were recognized at distance ranging from 15 to 150 m from the orchard edge.
Fig. 1 - Satellite image (a) and schematic diagram (b) of the P. alba’s clonal orchard located at the Estación Experimental Fernández (Convenio Universidad Católica de Santiago del Estero, Argentina). (a): Red symbols represent 47 external P. alba trees. Yellow lines delimit orchard borders. (b): T denotes seed controls that were removed in 2011. Grey and yellow areas represent failures and evaluated families (A-H), respectively.
At sampling, all trees were 13 years old. Adult fresh leaflets were taken from each tree and stored in a bag with silica gel until DNA extraction. One of the clones that was represented in all blocks was chosen randomly for seed analysis (clone 4). For adult genotyping, fresh leaflets were sampled from ramets at block VIII for all the clones but clone 5 that was sampled from block VII (Fig. 1) as it was missing in the former. Pods samples were collected from all ramets of the clone 4 located at all the different blocks (I-VIII). The progeny sample analyzed from of each mother plant (ramet) included eight seeds randomly taken from different pods, yielding 64 individuals.
DNA isolation and SSR analysis
Total genomic DNA was extracted using the DNeasy Plant® kit (Quiagen, Valencia, CA, USA) following the manufacturer’s instructions from leaves and 2- to 5-day-old seedling cotyledons. A total of 64 seeds and the 12 adult samples were characterized. Samples were genotyped at 10 SSR markers (Mo08, Mo09, GL6, GL8, GL9, GL12, GL15, GL18, GL21 and GL24) previously described for Prosopis ([34], [4]). PCR ampliïfications are described in Bessega et al. ([2], [5]) using FAM- or HEX fluorescent dye-label primers in a VERITI® termocycler (Applied Biosystems, Foster City, CA, USA). PCR products were electrophoresed by MACROGEN service (⇒ https://dna.macrogen.com/eng/index.jsp).
Data analysis
PCR products were sized using GeneMarker v. 1.91 ([40]). Genetic diversity was quantified through the number of alleles (A), observed (HO) and expected Nei’s diversity (HE), and polymorphic information content (PIC) indices, using the software Cervus ver. 3.0.7 ([33]).
In order to determine the power of the loci analyzed to discriminate between unique individuals we conducted two analyses. First, we obtained the genotype accumulation curve using the command “genotype_curve” of the “poppr” package ([27], [28]) in R ver. 3.5 ([38]), setting the number of resampling to 1000. Then, the accumulated probability of identity (PI) and the probability of identity when related individuals are included in the sample (PIsibs) were estimated using the software package GenAlex ver. 6.5 ([36]).
Parental analysis was performed following two approaches: (i) maximum likelihood, using the software Cervus ver. 3.0.7; and (I) Bayesian inference, using the MasterBayes package of R. Both methods are capable of incorporating missing data, genotyping errors and null alleles in parentage inference ([21], [26]).
Cervus
Paternity inference with Cervus ver. 3.0 assigns paternity using a statistics (delta), which is defined as the difference in the log-likelihood values of the two most likely pollen donors. In this sense, an individual will be assigned paternity if its likelihood is sufficiently higher than that of the second most likely male parent. Simulations (100.000) were run in order to estimate a critical value of delta above which a stated proportion of assignments in the population would be correct ([33], [26]). For this purpose, allele frequencies at each locus were estimated taking into account the unbalanced state of the orchard, and the parameters for simulating genotypes were: simulated offspring = 100.000; candidate parents = 12; proportion of candidate parents sampled = 0.6; proportion of loci typed = 0.91; proportion of loci mistyped = 0.01; and minimum typed loci = 8. Relaxed (80%) and strict (95%) confidence criteria were considered to assign paternity, following the manual recommendations ([33]).
MasterBayes
Based on genotypic information, the marginal posterior distributions of father-offspring pairs were used to calculate individual confidence levels in each assignment. This model was equivalent to the model run in Cervus except that confidence in relationships was assessed at the individual as opposed to the population level. First, the allelic frequencies were estimated considering the unbalanced design using the function “extractA”. Then, the offspring specific design matrices were constructed with the “varPed” function, assuming outcrossing = 1 based on Bessega et al. ([3]) and considering that G4 was the corresponding mother tree. Based on the Bayesian approach (using the “MCMCped” function), a categorical pedigree was generated in which only assignments with at least 80% confidence were accepted. This threshold was used for all MasterBayes assignments to allow comparison with relaxed confidence criteria used in Cervus analysis (see above).
Based on the resulting assignments, pollen contamination rate (m) in the clonal orchard, explained by the pollen contribution from the surrounding natural population, was estimated as the proportion of seeds which a father from the orchard could not be assigned. The individuals were considered as selfed when the assigned father genotype was the same as the known mother genotype. When the orchard seeds resulted from fertilization by pollen from outer trees, the prediction genetic gain is considered to be one half of that expected under no contamination ([41]). The expected reduction in genetic gain under contamination (GR) is therefore (eqn. 1):
where G is the gain expected under no contamination (taken as unity), assuming that external pollen sourced originated only from completely unimproved trees.
The paternity patterns were analyzed in different ways. First, the relationship among paternity and ramet abundance was evaluated using a generalized linear model with negative binomial distribution, with the “MASS” package ([44]) in R. This analysis was performed excluding G4, because according to Bessega et al. ([3]) selfing in P. alba is absent. Second, in order to quantify the effect of unequal male contribution, the status number (NS) was estimated by the following formula ([30] - eqn. 2)
where Σpi2 is the sum of the squared male contribution (pi). Finally, for each family (mother plant G4, individuals G4A to G4H), the minimal number of different fathers retrieved were counted and compared.
Results
The SSR loci analyzed were highly variables, showing between 2 and 12 alleles each (mean A value = 6.7 - Tab. 1). Averaged observed heterozygosity was higher than the expected heterozygosity. All the estimated genetic parameters indicated a rather high variability for the loci studied, and only 2 out of 10 loci (GL9 and MO09) showed a relatively low polymorphism (PIC < 0.25). The information produced by the 10 loci allowed the identification of 74 distinct multilocus genotypes in the clonal orchard, 10 corresponding to the adult trees (G1-G10) and 64 from the seeds analyzed.
Tab. 1 - Genetic diversity estimates of P. alba clonal orchard and details of the SSR loci analyzed. (Ta): Annealing primer temperature; (N): sample size; (A): number of alleles; (Ho, HE): observed and expected heterozygosity, respectively; (PIC): polymorphic information content; (fnull): null allele estimated frequency.
| Locus | Sequences (5′ -3′) | Ta (°C) | N | A | HO | HE | PIC | f null |
|---|---|---|---|---|---|---|---|---|
| GL6 | F: CTGGTTGCTGTGATTGGAGG R: CTCCAGGGATCACAAGACAAAC |
62 | 136 | 4 | 0.36 | 0.32 | 0.29 | -0.08 |
| GL8 | F: CAGGTGGGCATGAAGTTTCC R: CCAAGAACAACCTGCCGAAG |
58 | 131 | 12 | 0.80 | 0.74 | 0.72 | -0.04 |
| GL9 | F: ACTCTGCGGGTTAGGTAAGC R: ACCTGGAGCTGACATGGATC |
58 | 125 | 2 | 0.17 | 0.19 | 0.17 | 0.07 |
| GL12 | F: GAGTGAAGGTCGGGAAGAGG R: CCATTGGACCAAGGCAGAAC |
58 | 134 | 9 | 0.66 | 0.74 | 0.71 | 0.06 |
| GL15 | F: GTGTTATGGTCCCAACAGCC R: TGAAGAGGGAGGAATCGCAG |
58 | 127 | 8 | 0.80 | 0.78 | 0.75 | -0.01 |
| GL18 | F: GAGAATCTGGAGCAGCAACG R: AAGGTAGCGTCCCAGGTATG |
58 | 133 | 10 | 0.96 | 0.81 | 0.78 | -0.10 |
| GL21 | F: ATCTCCGTCACAACTTGCAC R: ACCCTCACTCCCGAATGATG |
58 | 136 | 4 | 0.49 | 0.55 | 0.47 | 0.05 |
| GL24 | F: CCTTAATCTCCCTCTCGGCC R: AACCAGGCTCTGCAGAAATG |
58 | 130 | 11 | 0.92 | 0.78 | 0.75 | -0.08 |
| MO08 | F: TATCCTAAACGCCGGGCTAC R: TCCCATTCATGCATACTTAAACC |
59 | 136 | 4 | 0.38 | 0.34 | 0.32 | -0.08 |
| MO09 | F: ATTCCTCCCTCACATTTTGC R: CATTATGCCAGCCTTTGTTG |
59 | 134 | 3 | 0.29 | 0.26 | 0.24 | -0.07 |
| Mean | - | - | 132.2 | 6.7 | 0.58 | 0.55 | 0.52 | -0.03 |
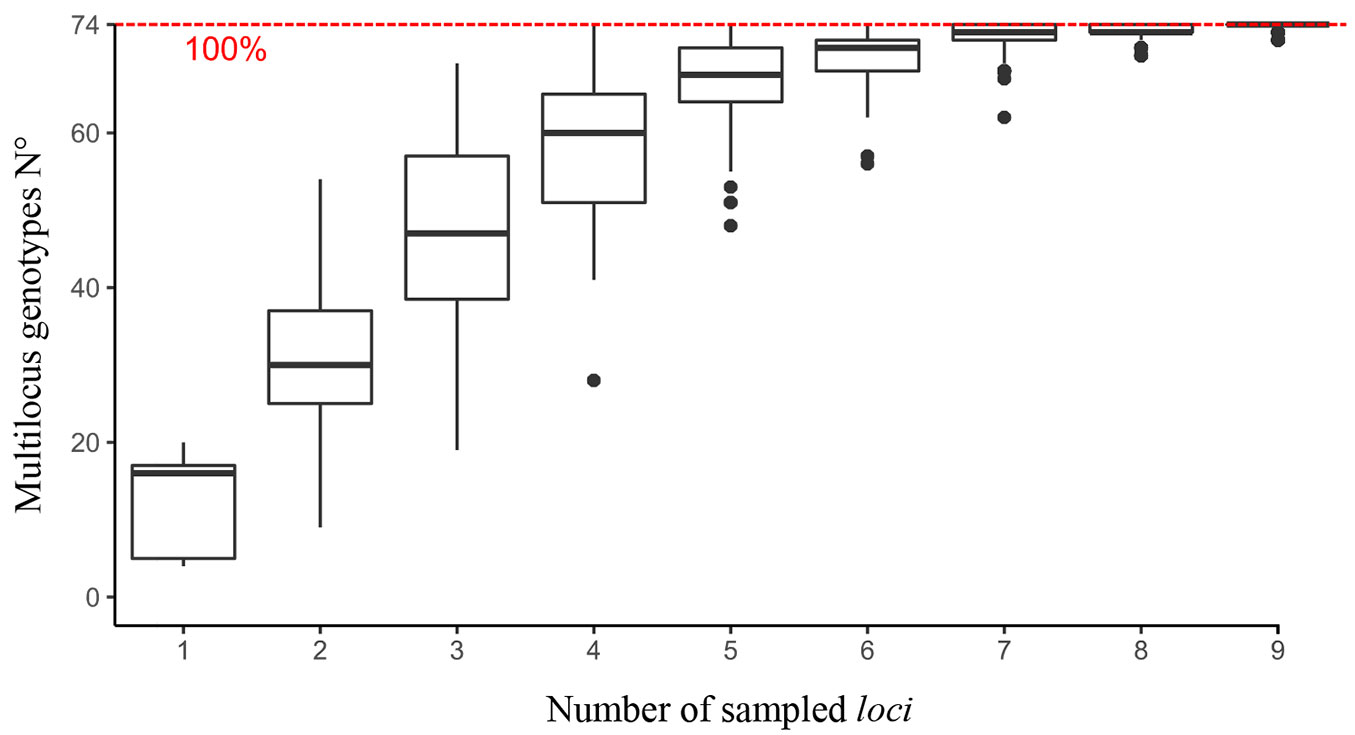
The analysis of the adult plants allowed to distinguish only 10 different multilocus genotypes, although the clonal orchard was established using 12 clones. Indeed, using the set of loci analyzed here clone 2 and clone 8 cannot be differentiated from each other and the same holds for clone 6 and clone 7. Each genotype can be cross referenced with the clonal selection made by Felker et al. ([18]) as follows. G1 corresponds to clone 1 (B2F17T2, block 2 family 17 tree 2 from R. Dulce), G2 to clone 2 (B6F12T1 - Rio Dulce) and clone 8 (B6F5T2 - Ibarreta), G3 to clone 3 (B1F8T4 - Castelli), G4 to clone 4 (B6F1T4 - R. Dulce), G5 to clone 5 ( B2F4T3 - Ibarreta), G6 to clone 6 (B1F5T3 - Pinto) and clone 7 (B1F6T3 - Castelli), G7 to clone 9 (B6F5T2 - Ibarreta), G8 to clone 10 (B5F9T2 - Castelli), G9 to clone 11 (B1F5T4 - Ibarreta) and G10 to clone 12 (B7F6T4 - Castelli). The genotype accumulation curve (Fig. 2) suggests that the full recognition (100%) of all the studied multilocus genotypes can be achieved using only 9 loci. The accuracy of the individual identification was high considering that the accumulated probability of identity (PI) and probability of identity when related individuals are included in the sample (PIsibs) were 7.2 · 10-7 and 0.003, respectively.
Fig. 2 - Genotype accumulation curve. Box plots were constructed from the number of observed genotypes considering a randomized bootstrap without resampling from the original data.
Paternity analysis
Null allelic frequencies were considered negligible as they were close to zero for most of the analyzed loci, being between 0.05 and 0.1 only in three cases (Tab. 1). Based on Cervus analysis, the average non-exclusion probability for one candidate parent given the genotype of the known mother was low (NE-2P= 0.006) and no incongruence was detected among progeny and the known mother genotypes. From the total offspring analyzed, 47 individuals (seeds) were successfully assigned to one parent (pollen donor) inside the clonal orchard, of which 44 using the strict criterion and three with the relaxed one (Tab. S1 in Supplementary material). These values indicate that 69-73% of the analyzed seeds have been pollinated by a donor from the orchard, according to the different confidence criteria. The maximum pollen contamination rate (m) was 31%. G4 was assigned as male parent in only one case, yielding a selfing rate of 0.016.
Based on the posterior probability of each parental assignment obtained using MasterBayes (Tab. S1 in Supplementary material), 63-72% of the offspring exhibited multilocus genotypes compatibles with pollen donors from the clonal orchard, considering strict and relaxed criteria, respectively. Consequently, the external pollen contamination rate was 28-37% and no selfing was detected. The analysis of seeds originated by external pollen donors indicated that the number of external trees acting as male parents was only 2.6 (CI95%= 1.2-5.0).
A high consistency was found (89% of cases based on relaxed criteria) between the parental assignments using both maximum likelihood and Bayesian approaches (see Tab. S1 in Supplementary material). In 67% of cases the consistent assignments were from internal pollen donors, while 22% were from external male parents. The remaining 11% of cases showed differences among algorithms: 6% were assigned only by Cervus and 5% only by MasterBayes.
The maximum pollen contamination rates estimated by both methods allowed to assess a decrease in the expected clonal orchard genetic gain from 15.5 to 18.5%. All the genotypes, with the exception of G1, were able to fertilize the G4 ramet egg cells. G6 was the clone that most contributed to G4 offspring (Tab. S1 in Supplementary material).
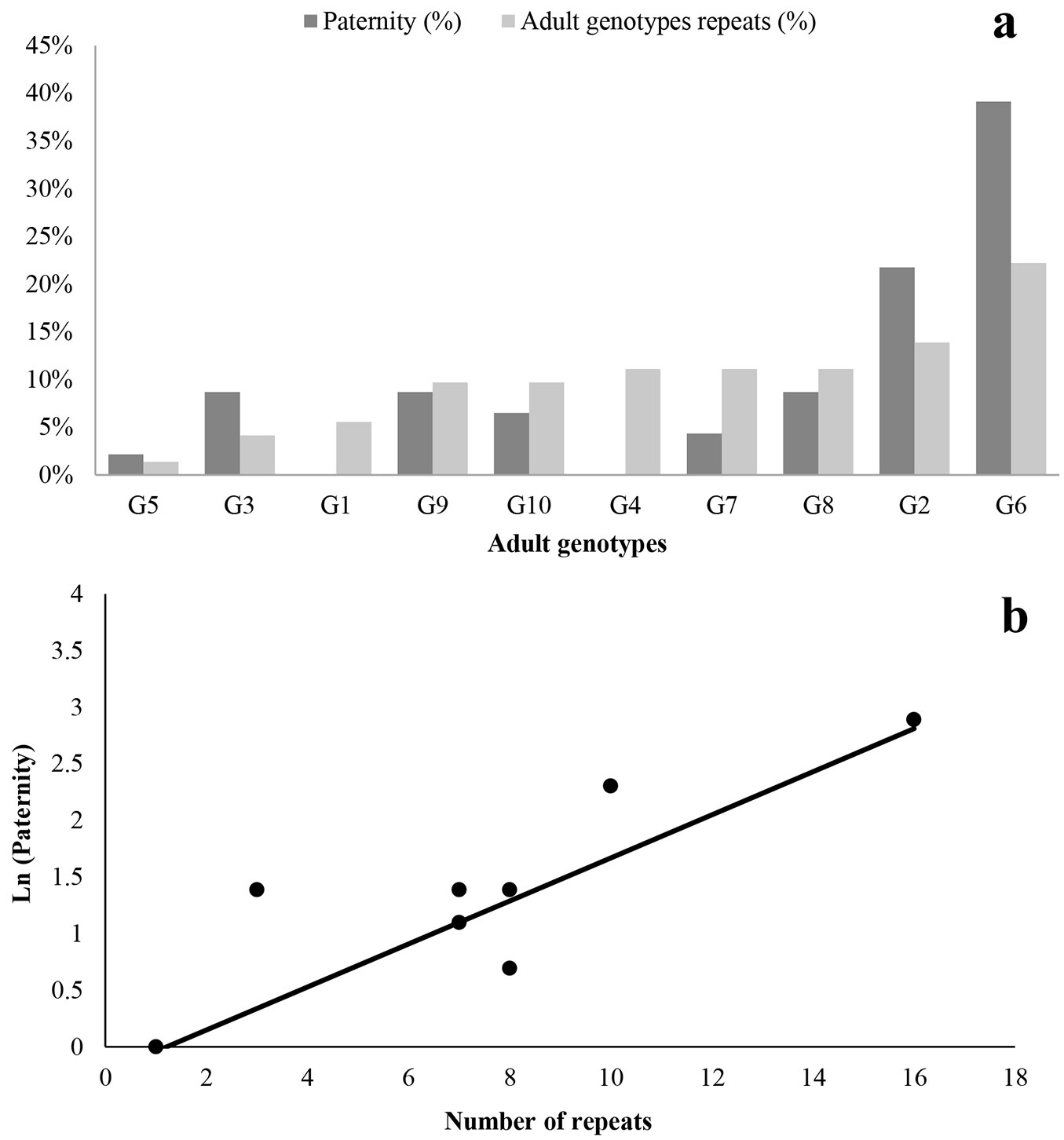
The number of offspring assigned to each clone varied among 0 to 18 (Tab. 2). The different contribution of different clones to G4 offspring may be partially explained by the variation in the number of surviving ramets. Despite the orchard had originally been designed with a balanced design, the rooting of the cuttings was not equally effective. Therefore, 32 trees were not present at the time of sampling, which caused the design to be treated as unbalanced (Fig. 1b, Fig. 3a). At sampling, only one out of 8 ramets corresponding to G5 was present, but G6 exhibited 16 repeats as it represents clones 6 and 7. The regression of paternity success on the number of surviving ramets of each clone (Fig. 3b) is positive and highly significant (P= 0.0004). The status number (Ns), based on the relative contribution of each genotype in the gene pool of male gametes (Tab. 2), was 4.4 and the minimum number of different male parents that pollinated each mother plant (family G4, individual G4A to individual G4H) varied from 3 to 5, with an average of 4.4 (Tab. 2).
Tab. 2 - Number of seed assigned to each genotype, considering the different evaluated families (mother tree G4A to mother tree G4H). dN represents the number of descendants corresponding to each adult genotype; pi represents the relative contribution of each genotype over the total evaluated progeny; ri represents the proportion of clone repeats in the orchard at sampling.
| Assigned Male Parent |
Families | dN | p i | r i | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| G4A | G4B | G4C | G4D | G4E | G4F | G4G | G4H | ||||
| G1 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.00 | 0.06 |
| G2 | 1 | 0 | 0 | 1 | 3 | 2 | 3 | 0 | 10 | 0.16 | 0.14 |
| G3 | 1 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 4 | 0.06 | 0.04 |
| G4 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.00 | 0.11 |
| G5 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0.02 | 0.01 |
| G6 | 2 | 4 | 1 | 3 | 2 | 1 | 0 | 5 | 18 | 0.28 | 0.22 |
| G7 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 0 | 2 | 0.03 | 0.11 |
| G8 | 0 | 0 | 1 | 1 | 0 | 1 | 0 | 1 | 4 | 0.06 | 0.11 |
| G9 | 0 | 0 | 1 | 0 | 0 | 1 | 2 | 0 | 4 | 0.06 | 0.10 |
| G10 | 0 | 1 | 2 | 0 | 0 | 0 | 0 | 0 | 3 | 0.05 | 0.10 |
| External | 4 | 0 | 3 | 3 | 3 | 3 | 1 | 1 | 18 | - | - |
| Pollen donors | 4 | 5 | 5 | 4 | 3 | 5 | 5 | 4 | - | - | - |
| Mean | 4.4 | - | - | - | |||||||
Fig. 3 - (a) Paternity and representation for each adult genotype of the orchard. (b) Relationship among the number of descendants produced by each genotype (Ln paternity) and repeats in the orchard. Lineal regression (y = 0.19x - 0.23) for a GLM distribution considering a negative binomial distribution and overdispersion Φ = 1.34.
Discussion
The establishment of seed clonal orchards is an important step of the Prosopis alba breeding programmes in Argentina. In order to provide valuable information for the development of these orchards and suggest future management strategies, we estimated the pollen contamination rate and pollination patterns in a clonal P. alba stand located in Santiago del Estero, Argentina.
SSR markers are largely used tools for parental analyses ([43], [19], [48]). The SSR loci used here were highly polymorphic and null allele frequencies were negligible for all loci. The high genetic variability detected allowed to discriminate individual genotypes and to exclude unrelated candidate trees from parentage of any arbitrary offspring. Moreover, the genotype accumulation curve suggests that the combination of markers used were adequate to discriminate all individuals of the progeny. The lack of differentiation between some pairs of clones showing the same multilocus genotype could be solved by analyzing a larger number of molecular markers both in the clones and the corresponding plants they originated from.
Pollen contamination is an important factor that reduces the expected genetic gain in a clonal orchard. We demonstrated that between 27% and 37% of the sampled seeds are not genetically compatible with male parents located inside the clonal orchard, according to the different estimation methods and confidence criteria. These contamination values are within the range recorded in the literature for both wind and insect pollinated orchards. Bilgen & Kaya ([8]) reported that 29% of the embryos of Pinus brutia were sired by pollen sources outside the orchard. Feng et al. ([19]), Torimaru et al. ([43]) and Buiteveld et al. ([9]) estimated contamination rates of 25%, 51% and 70% for Pinus koraiensis, Pinus silvestris, and Quercus robus, respectively. In Juglans nigra, Ebrahimi et al. ([14]) proved that the within-orchard pollination was significantly higher in an isolated orchard than in a non-isolated one. For the insect pollinated Schima superba, Yang et al. ([48]) detected a lower, but important, level of pollen contamination (7.01%) originated from outside the orchard. In all these cases, individuals of the same species or a close related one could be found near the clonal orchard.
Failures in the recognition of parental trees within the studied orchard can be explained by: (i) natural regeneration of individuals from outside (e.g., seeds carrying unrelated genotypes) establishing within the clonal orchard; (ii) mutational events (such as inversions) in the offspring, whose rate is fairly high in SSR regions of the genome; (iii) scoring mistakes; and (iv) pollen flow from outside the orchard ([37], [8]). Some of these causes can be easily ruled out in our case. First, natural regeneration from outside could be excluded as the orchard is controlled by constant thinning in order to preserve the genotypes of interest. Secondly, mutational events can be considered as unlikely because no mismatch was found between seed multilocus genotypes and the known mother genotype. Genotyping errors might skew the estimation of pollen contamination, but the error quantification estimated by MasterBayes based on the 10 SSR loci analyzed was very low (being the allelic drop out and stochastic error equal to 0.002 and 0.02, respectively).
Considering that other P. alba trees were found near around the orchard, external pollen contamination could be the most likely reason to explain the occurrence of multilocus genotypes not attributable to male parents among the clones that constitute the trial. Pollen contamination rates here obtained can be explained by the contribution of 1 to 5 different trees from outside the orchard that were acting as pollen donors. A low number of parents participating in pollination was also observed in natural populations of insect-pollinated tree species ([3], [35]). The level of pollen contamination depends on factors like the amount of pollen produced inside the orchard, flowering synchronization among the clones, and annual weather variation ([24], [10]). It is not possible to determine which factor mostly affected the results of this study. Asynchronous reproductive phenology has been reported to cause temporal isolation between the orchard’s breeding subpopulations of Douglas fir, leading to increased pollen contamination and increased selfing (due to the decreased number of pollen donors and to the reduction of gametic competition between orchard and outside parents) increasing the chance of gene flow ([15]). The non-synchronous flowering among families was reported as the major cause for differences among the pollen clouds of each family in Schima superba ([48]), where precocity was described for some families. Additionally, the variation of climatic factors, such as rain, frost and prevalent winds, may represent a gene flow barrier. Indeed, rainfall may strongly affect flowering and/or pollination, and intensive raining episodes may cause different clones to be differentially affected, yielding differential amount of pollen. Furthermore, insects like Prosopidopsylla flava and fungal pathogens like Pestalotiopsis spp. are known to attack mature and immature leaves and flowers of Prosopis, and the Argentine parrots (Myiopsitta monachus) to feed on clone pods. As some clones in the studied orchard are more precocious than others, it may be hypothesized that they could represent traps or escapes to pests’ invasions. Finally, the entomophilous pollination may be also affected by rainy periods, when the insects’ visiting rate of flowers is reduced, thereby differently influencing the pollination of clones with different blooming time. The lack of studies on phenology and pollen production in the studied orchard is a limitation of the present investigation that can help clarifying the aforementioned issues. The importance of gaining greater empirical knowledge of factors affecting pollen-mediated gene flow from Prosopis species is needed for avoiding gain loss. To maximize the genetic gain, this kind of studies must be repeated at different times to quantify possible variations in pollen contamination rate or, alternatively, establish seed orchards in areas well isolated from other pollen sources.
Pollen contamination from the wild to the orchard is relevant in terms of the possible reduction of the expected gains. On the other hand, pollen from the orchard (especially in case of exotic material) could represent a harmful contamination of natural populations in the surrounding environment. The risk of gene escape from the orchard to the wild may be evaluated by fine scale genetic analyses, flowering synchrony, analysis of barrier at pre- and post-pollination and post-seed dispersal, as proposed by Barbour et al. ([1]) in Eucalyptus species. The studied orchard has been established not far from other Prosopis stands near the experimental station. Avoiding contamination from any other sources is crucial for setting up new areas of seed collection to be cataloged as selected material. The results of this study are relevant for future implementations of Prosopis orchards for seed production, since the current stands include materials from different distant sources.
Parental inference was carried out using both likelihood and Bayesian approaches. Here, such methodologies mainly differ in the algorithm used to estimate the confidence of each assignment. While Cervus uses population-level confidence, MasterBayes relies on individual-level confidence thresholds. Nonetheless, the results obtained by both approaches were highly consistent. Similar results have been reported by Walling et al. ([45]), who found only 7 cases of parental assignment mismatches between the two methods. However, in all cases the main difference was attributable to different confidence levels, as both approaches indicated the same parental multilocus genotypes as the most probable parents.
The unequal contribution of different parental clones in the orchard to the studied offspring yielded a status number (Ns) of 4.4, i.e., the expected coancestry in the seed crop is equal to a Mendelian population with an effective size of 4-5 individuals. Hansen & Kjaer ([22]) found a Ns= 4.2 in a Abies nordmanniana orchard including 13 different clones, while Hansen ([23]) estimated Ns= 4.6 for a A. alba orchard with 12 different clones. It was reported that the number of successful pollinations by each clone is highly related to its number of ramets in the orchard ([14]). Populations with low effective sizes may increase the level of relatedness between seedlings in the orchard, leading to genetic drift and inbreeding depression in future crosses ([47]). In addition, the estimation of a status number lower than expected may indirectly increase the selective process that is done during the constitution of the orchard.
In this study the estimated number of pollen donors per mother plant was near four. This result is fairly similar to the findings of Bessega et al. ([3]) in a P. alba natural population, where six pollen donors per mother tree were assessed by indirect methods. Moreover, the low selfing rate obtained in this study is in agreement with previous studies on this species in the wild ([3]). It may be argued that the presence of multiple ramets of the same clone within the orchard led to an increased level of selfing. However, the low proportion of selfed offspring in the orchard is not a relevant problem from the productive point of view, as reported for other forest species ([29], [23], [43]). Our results were also consistent with previous studies on the insect-pollinated Schima superba ([48]), where a small number of effective pollen donors per mother tree (2.3) and a high overall crossover rate (98.5%) were also reported. According to Yang et al. ([48]), it can be suggested that pollen distance and dispersion in trees pollinated by bees (as is the case of P. alba) depends on several biotic and abiotic factors such as the distance between individuals, phenology of flowering, behavior of the dispersing agent and weather conditions.
Conclusion
The Prosopis alba orchard analyzed in this study is the only one established by vegetative propagation for species of the genus Prosopis. The lack of similar trials is due to technical difficulties to obtain rooted cuttings able to successfully develop in the field. Despite the limitations due to the unbalanced design and the relatively low number of clones currently available, our results support the usefulness of genetic analyses in forest breeding and management programmes, particularly for the establishment and monitoring of trials. A recommended management strategy to reduce pollen contamination should consider the establishment of seed orchards in areas well-isolated from putative sources of contamination. Moreover, the removal of related trees acting as non-selected pollen donors or the use of gene flow barriers is encouraged. Finally, the equal representation of genotypes within the orchard is crucial to guarantee similar contributions of each genotype in the seed production. The lack of descendants for some genotypes observed in this investigation highlights the importance of conducting preliminary studies on synchrony and compatibility among the selected genotypes.
Acknowledgements
This research was supported by funding from Agencia Nacional de Promociones Científicas y Técnológicas (ANPCyT) PICT 2013-0478, PICT2016-0388 and Universidad de Buenos Aires (UBA): 20020170100093BA given to JVC, CP and CB.
References
Gscholar
Gscholar
Gscholar
Gscholar
CrossRef | Gscholar
Gscholar
Gscholar
Supplementary Material
Authors’ Info
Authors’ Affiliation
Cesar Vilardi Juan
Ofelia Saidman Beatriz 0000-0003-4704-3713
Cecilia Bessega 0000-0002-8575-1828
Departamento de Ecología, Genética y Evolución, Facultad de Ciencias Exactas y Naturales, Universidad de Buenos Aires (Argentina)
Ofelia Saidman Beatriz 0000-0003-4704-3713
Cecilia Bessega 0000-0002-8575-1828
Instituto de Ecología, Genética y Evolución (IEGEBA) CONICET-UBA, Buenos Aires (Argentina)
Estación experimental Fernández, Universidad Católica de Santiago del Estero (UCSE), Santiago del Estero (Argentina)
Corresponding author
Paper Info
Citation
D’Amico I, Vilardi Juan C, Saidman Beatriz O, Ewens M, Bessega C (2019). Pollen contamination and mating patterns in a Prosopis alba clonal orchard: impact on seed orchards establishment. iForest 12: 330-337. - doi: 10.3832/ifor2936-012
Academic Editor
Alberto Santini
Paper history
Received: Jul 31, 2018
Accepted: Apr 05, 2019
First online: Jun 14, 2019
Publication Date: Jun 30, 2019
Publication Time: 2.33 months
Copyright Information
© SISEF - The Italian Society of Silviculture and Forest Ecology 2019
Open Access
This article is distributed under the terms of the Creative Commons Attribution-Non Commercial 4.0 International (https://creativecommons.org/licenses/by-nc/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Web Metrics
Breakdown by View Type
Article Usage
Total Article Views: 45916
(from publication date up to now)
Breakdown by View Type
HTML Page Views: 37899
Abstract Page Views: 4015
PDF Downloads: 3169
Citation/Reference Downloads: 2
XML Downloads: 831
Web Metrics
Days since publication: 2532
Overall contacts: 45916
Avg. contacts per week: 126.94
Article Citations
Article citations are based on data periodically collected from the Clarivate Web of Science web site
(last update: Mar 2025)
Total number of cites (since 2019): 1
Average cites per year: 0.14
Publication Metrics
by Dimensions ©
Articles citing this article
List of the papers citing this article based on CrossRef Cited-by.
Related Contents
iForest Similar Articles
Review Papers
Perspective on the control of invasive mesquite trees and possible alternative uses
vol. 11, pp. 577-585 (online: 25 September 2018)
Short Communications
Towards an optimal sampling effort for paternity analysis in forest trees: what do the raw numbers tell us?
vol. 5, pp. 18-25 (online: 27 February 2012)
Technical Reports
Conservation and use of elm genetic resources in France: results and perspectives
vol. 13, pp. 41-47 (online: 03 February 2020)
Research Articles
Delineation of seed collection zones based on environmental and genetic characteristics for Quercus suber L. in Sardinia, Italy
vol. 11, pp. 651-659 (online: 04 October 2018)
Research Articles
Genetic analysis of Latvian Salix alba L. and hybrid populations using nuclear and chloroplast DNA markers
vol. 10, pp. 422-429 (online: 24 March 2017)
Research Articles
The maternal environment of European beech (Fagus sylvatica L.) affects intrapopulation variability in seed traits and germination
vol. 19, pp. 114-121 (online: 10 April 2026)
Research Articles
Seed germination traits of Pinus heldreichii in two Greek populations and implications for conservation
vol. 15, pp. 331-338 (online: 24 August 2022)
Research Articles
Impact of inbreeding on growth and development of young open-pollinated progeny of Eucalyptus globulus
vol. 15, pp. 356-362 (online: 20 September 2022)
Research Articles
Patterns of genetic variation in bud flushing of Abies alba populations
vol. 11, pp. 284-290 (online: 13 April 2018)
Research Articles
Networking sampling of Araucaria araucana (Mol.) K. Koch in Chile and the bordering zone of Argentina: implications for the genetic resources and the sustainable management
vol. 2, pp. 207-212 (online: 22 December 2009)
iForest Database Search
Search By Author
Search By Keyword
Google Scholar Search
Citing Articles
Search By Author
Search By Keywords
PubMed Search
Search By Author
Search By Keyword