Genetic analysis of Latvian Salix alba L. and hybrid populations using nuclear and chloroplast DNA markers
iForest - Biogeosciences and Forestry, Volume 10, Issue 2, Pages 422-429 (2017)
doi: https://doi.org/10.3832/ifor2004-009
Published: Mar 24, 2017 - Copyright © 2017 SISEF
Research Articles
Abstract
Latvia is at the northern border of the species range of S. alba L. in Europe, and there has been some debate whether the Latvian populations of S. alba are autochthonous, as this species has long been planted in Latvia as an ornamental tree in gardens, parks and roadsides. In addition, there is increasing interest in the use of several Salix species (including S. alba) as bioenergy crops. Natural S. alba stands throughout Latvia, as well as stands of possibly hybrid origin were analysed using nuclear and chloroplast markers. Our results showed that S. alba populations are probably natural, and that the rate of vegetative reproduction is low, supporting the evidence that Latvia is within the natural range of S. alba. These results provide the basis for the identification of possibly introduced or artificially regenerated stands of S.alba in Latvia. In addition, our results confirm that S. alba hybridises with S. fragilis, and that natural stands including hybrid individuals can be established. The analysis of chloroplast markers indicated that the predominant hybridisation occurs by fertilisation of S. fragilis by S. alba pollen; however, the extent of haplotype sharing between these two species should be further investigated.
Keywords
Introduction
There is an increasing interest of using short rotation species such as white willow (Salix alba L.) as a source of biomass in Latvia ([33]). Currently, foreign clones of this species are mainly utilised in plantations, though there are plans to identify high-yield locally adapted clones. To this purpose, information about the structure, provenance and diversity of Salix alba populations in Latvia will assist in the identification of the most valuable genetic resources for further utilisation.
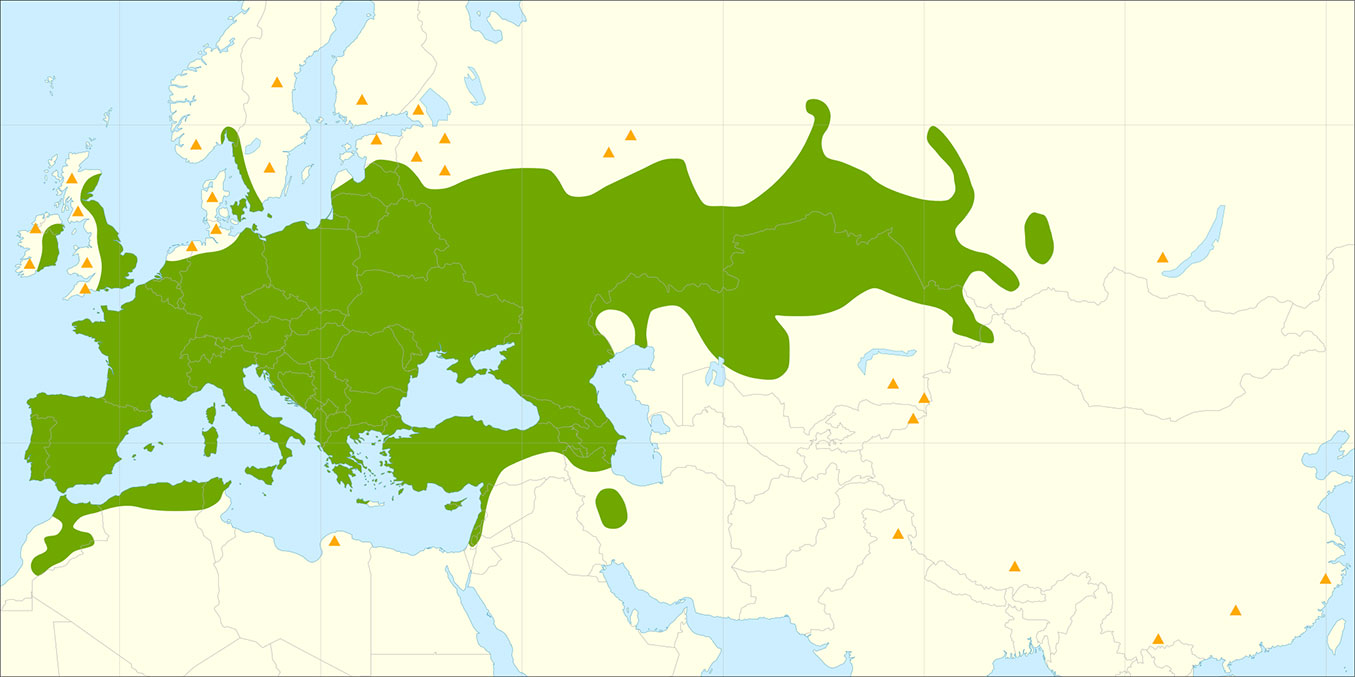
In Latvia, as well as Estonia and Lithuania, white willow has long been planted as an ornamental tree in gardens, parks and roadsides. White willow has intensively spread into surrounding areas due to the high reproductive capacity of this species, both sexual and vegetative. As a consequence, different opinions exist among researchers about whether this species is introduced or if it is native. The status of this species in Latvia is further complicated by two other factors. Firstly, white willow has many sub-species and varieties, and also readily forms hybrids with other Salix species. Cinovskis et al. ([6]) consider these sub-species and varieties as cultivated clones or decorative forms, which should not be considered at a taxonomic level. Secondly, various studies consider that Latvia is at the northern range boundary (or possibly outside) of the natural distribution area of this species, and therefore the provenance of particular white willow individuals or populations may be difficult to determine. Several studies ([24], [39]) place the distribution boundary of white willow to the north of Latvia, extending from lake Onega in Russia along the Svir River to lake Ladoga, then along the Neva River, the Gulf of Finland, and the east coast of the Baltic Sea, thus encompassing all of the Baltic States (Estonia, Latvia and Lithuania - Fig. 1). In contrast, other authors considered the boundary to be in northern Latvia, based on the extensive natural white willow stands found along the Gauja River ([34]). This northern boundary along the Latvian-Estonian border was also reported in the USSR Atlas of Woody Plants ([35]), as well as in the European Flora Atlas, where S. alba stands in Latvia and Lithuania are reported as autochthonous, while Estonian stands are considered to be of unknown provenance ([15]). The autochthonous status of white willow stands along the Gauja and Daugava rivers has been emphasised by Latvian dendrologists ([21], [9]). Evidence supporting this conclusion includes the presence of white willow in mixed species stands (including Alnus incana, Alnus glutinosa, Ulmus glabra and other species), as well as the mixed age structure of white willow stands ([8], [20]), while white willow is considered to form only even-aged stands at the northern distribution boundary ([21], [18], [35]). Other reports suggest that the northern distribution boundary might be in the southern part of the Baltic states, with mixed age natural stands of white willow found only along the Svete River in Latvia, and in particular along the Nemuna, Neri and Neveži rivers in Lithuania ([5], [22], [25]).
Fig. 1 - Natural distribution range of white willow (Salix alba L. - in green). Orange triangles represent putative introduced and/or naturalized (synanthropic) populations. Source: Houston Durrant et al. ([14]).
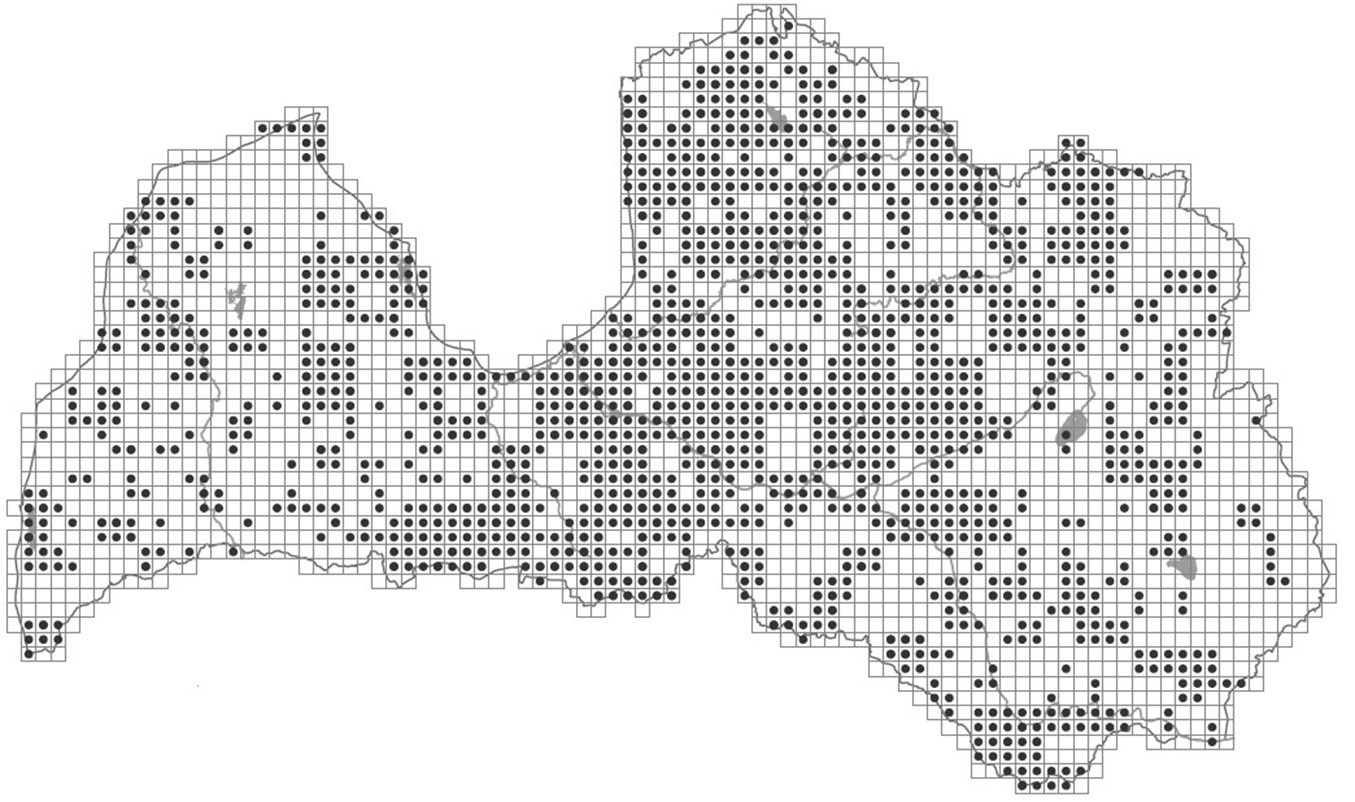
Despite these uncertainties about the provenance and distribution area of white willow in the Baltic States, this species is found throughout this region, and was mentioned in the first description of the flora of the region ([10]), as well as in further descriptions of the flora of the Baltic region. In the 19th century, Salix alba was reported to be a widespread species in the eastern Baltic region, commonly found in natural stands along sandy riversides and moist forests, as well as in artificial plantations ([17]). In the Atlas of Latvian Woody Plants, Salix alba is reported in 1349 sites as the dominant species (in both plantations and natural stands, grid size 5×5 km, frequency 48 %; [20] - Fig. 2), and is the most common Salix species in Latvian parks and plantings ([4]). Many S. alba cultivars have been cultivated for long time: cultivar ’Calva’ was first described in 1700, ’Chermesina’ and ’Sericea’ approximately in 1840 ([19]). The variety or cultivar ’Vitellina’ was described as a separate species by Linnaeus in 1671 (S. vitellina L. - [9]), but the taxonomic status of these cultivars, sub-species or species is currently disputed ([5]). In this study, the samples collected from the National Botanic Garden (Salaspils, Latvia) are designated as cultivated clones or varieties. These cultivated S. alba varieties were compared with the putative natural populations of S. alba in Latvia to determine the genetic similarity between the cultivated and natural forms, and to possibly identify the escape of cultivated clones into natural habitats.
Fig. 2 - Distribution of white willow (Salix alba L.) in Latvia. Source: Laivinš et al. ([20]).
Various kinds of DNA markers have been used to analyse Salix species, including dominant markers such as AFLPs, ISSR and RAPDs ([23]), as well as co-dominant markers such as SSRs ([30]). Salix species have also been extensively studied with maternallyinherited chloroplast markers ([28], [30]), which enables the differentiation of lineages, as well as the direction of hybridisation. S. alba and S. fragilis are both polyploids, with S. alba reported as being stably tetraploid ([11]), and S. fragilis being mainly tetraploid, but with other ploidy levels also identified ([1]). SSR markers have been developed for a range of Salix species, and many SSR markers showed cross-species amplification in other Salix species and genera such as Populus ([3], [13]). Analysis of polyploid species such as S. alba and S. fragilis with codominant markers such as SSRs is complicated by the difficulty of determining the allele dosage, and therefore they are often converted to binary, dominant marker genotypes for analysis. However, some Salix SSR markers have been reported to have disomic-monogenic inheritance in tetraploid S. alba and S. fragilis, and therefore can be analysed as diploid, codominant markers ([16]). Identification of such SSR markers is time-consuming as the segregation of alleles must be determined in the progeny of controlled crosses.
There is evidence that interspecific hybridisation is common between a range of Salix species ([28]), and controlled crosses have shown that S. alba and S. fragilis can also readily hybridise ([37]). Additionally, continuous variation of morphological characteristics between these species has been reported in natural populations ([38]). Despite this morphological continuum between the two species, DNA marker studies have indicated that the two species are clearly differentiated, suggesting that introgressive hybridisation is not widespread ([38], [27]).
Given the increasing interest in Latvia of utilising S. alba as a short-rotation plantation species, we used nuclear and chloroplast markers to analyse the genetic diversity and population structure of natural S. alba stands throughout Latvia, with the aim of characterising the available genetic resources for this species in this region. In addition, stands of S. fragilis and putative hybrid individuals were analysed to investigate hybridisation between these two species. As the autochthony of Latvian S. alba populations is uncertain, individuals representing cultivated putative subspecies and other decorative forms were analysed, to determine their genetic similarity to natural S. alba stands in Latvia.
Materials and methods
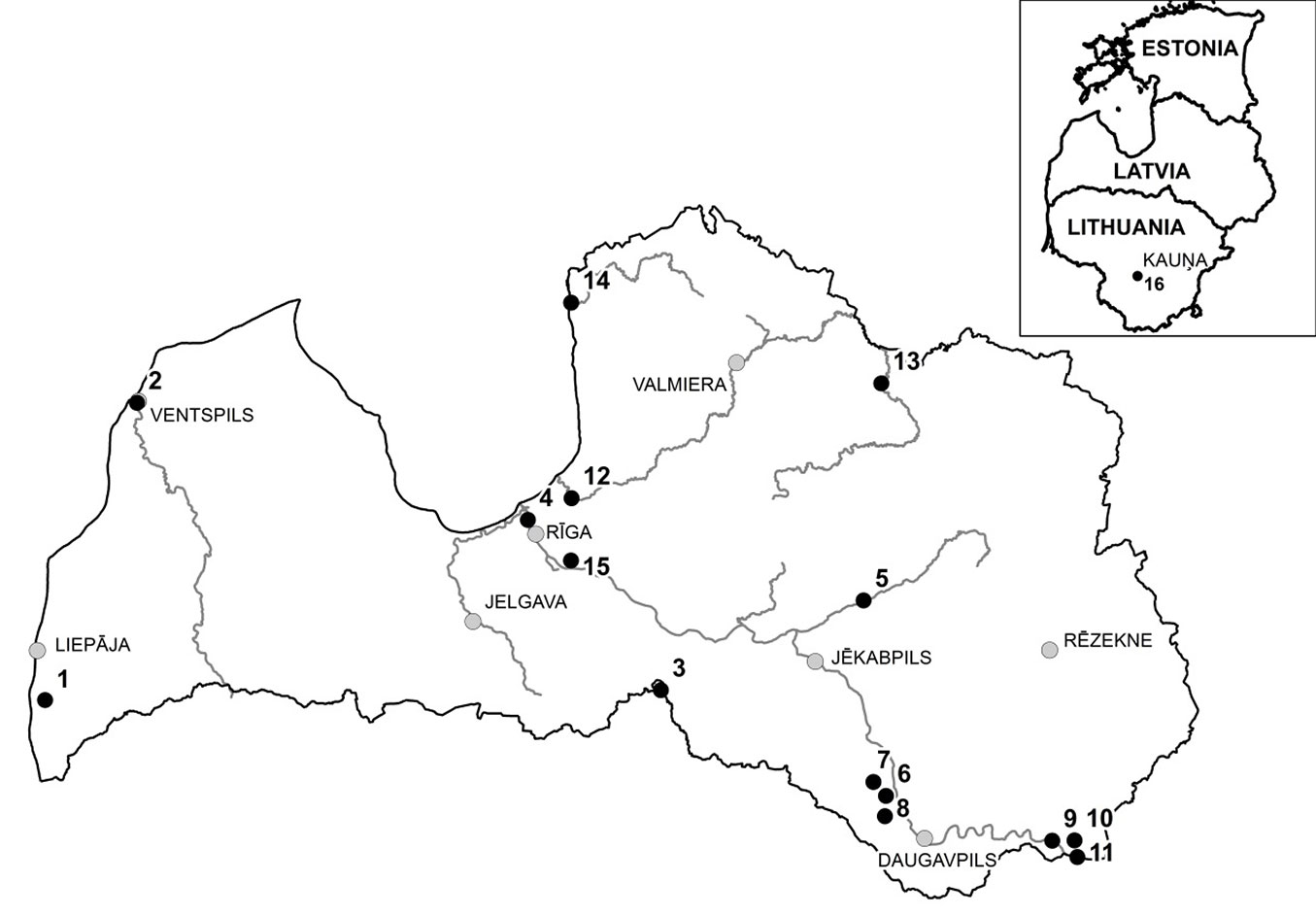
Leaf samples of S. alba and putative hybrids with S. fragilis (based on phenotypic traits) were collected from 14 stands in Latvia, and one in Lithuania (Fig. 3, Tab. 1). Leaf morphological traits were analysed to differentiate the two species and to identify putative hybrids, particularly leaf colour/pubescence and serration. The populations were designated as S. alba, S. fragilis or hybrid populations, according to the dominant species, and the sampled individuals were morphologically selected according to the dominant species in each stand (Tab. 1). Individuals with hybrid morphology were dominant in only one population (Dviete). In addition, leaf samples from one individual of each of the S. alba cultivars ’Chermesina’, ’Britzenis’, ’Calva’, ’Coccinea’, ’Vitellina’, ’Sericea’, one individual of S. alba and two individuals of Salix × rubens Schrank (hybrid between S. alba and S. fragilis) were collected from the National Botanic Garden in Salaspils, Latvia (Tab. 1).
Fig. 3 - Location of sampled stands (black circles) in Latvia and Lithuania. Population numbers are the same as in Tab. 1 (see below).
Tab. 1 - Locations and description of sampled stands in Latvia and Lithuania.
| Pop. no. | Location | River catchment |
Coordinates | Species/ Cultivar |
Sample size |
|---|---|---|---|---|---|
| 1 | Nica | Barta | 56°21′; 21°04′ | S. alba | 24 |
| 2 | Ventspils | Venta | 57°23′; 21°34′ | S. alba | 26 |
| 3 | Vecmemele | Memele/Lielupe | 56°24′; 24°54′ | S. alba | 23 |
| 4 | Voleri/Riga | Daugava | 57°00′; 27°04′ | S. alba | 24 |
| 5 | Laudona | Aiviekste/Daugava | 56°42′; 26°11′ | S. alba | 24 |
| 6 | Dviete | Dviete/Daugava | 56°03′; 26°16′ | S. alba × S. fragilis | 15 |
| 7 | Bebrene | Dviete/Daugava | 56°04′; 26°12′ | S. alba | 7 |
| 8 | Ilukste | Daugava | 55°57′; 26°16′ | S. fragilis | 24 |
| 9 | Lielindrica | Daugava | 55°51′; 27°20′ | S. alba | 24 |
| 10 | Lielindrica | Daugava | 55°50′; 27°19′ | S. fragilis | 24 |
| 11 | Piedruja | Daugava | 55°47′; 27°26′ | S. fragilis | 24 |
| 12 | Adaži | Gauja | 57°04′; 24°21′ | S. alba | 18 |
| 13 | Vireši | Gauja | 57°27′; 26°20′ | S. alba | 27 |
| 14 | Salaca | Salaca | 57°45′; 24°21′ | S. alba | 21 |
| 15 | Salaspils/ National Botanic Garden |
Daugava | 56°51′; 24°21′ | ’Chermesina’ | 1 |
| ’Britzenis’ | 1 | ||||
| ’Calva’ | 1 | ||||
| ’Coccinea’ | 1 | ||||
| ’Vitellina’ | 1 | ||||
| ’Sericea’ | 1 | ||||
| S. alba | 1 | ||||
| Salix × rubens | 2 | ||||
| 16 | Kaunas | Nemuna | 54°54′; 23°53′ | S.alba | 17 |
DNA from leaves was isolated using a CTAB-based method ([32]). A total of 331 individuals were genotyped using eight nuclear SSR markers and six angiosperm chloroplast markers. Genotyping was carried out using nuclear SSR markers SB243, SB93, SB38, SB80, SB201, SB24, SB194 and SB199, which were reported to be informative in a range of Salix species ([3]). Six angiosperm chloroplast markers ccmp2, ccmp4, ccmp6, ccmp10, ccmp7 and ccmp3 were also utilised ([41]), which had previously been used to analyse S. caprea ([28], [30]). The PCR reactions were multiplexed as follows: set A (SB243, SB93, SB38); set B (SB80, SB201); set C (SB24, SB194, SB199); set D (ccmp2, ccmp3, ccmp10); set E (ccmp 4, ccmp6, ccmp7). Each forward primer in the multiplex sets was labelled with a different fluorophore (6-FAM, HEX or TMR) to facilitate visualisation using capillary electrophoresis. PCR reactions for the nuclear SSR markers were carried out in a 20 μl solution comprising: 4 μl of 5× HOT FIREPol® Blend Master Mix (containing 10 mM MgCl2, 2 mM dNTPs and BSA - Solis BioDyne, Estonia), 0.4 μM final concentration of each primer and 1 μl genomic DNA. The PCR profile consisted of an initial denaturation at 95 °C for 15 min, followed by 35 cycles of denaturation at 94 °C for 40 s, annealing at 48 °C for 60 s, and extension at 72 °C for 120 s. Final extension of amplified DNA occurred at 72 °C for 10 min. PCR reactions for the chloroplast SSR were carried out in a 10 μl solution comprising: 2 μl of 5× HOT FIREPol® Blend Master Mix containing 10 mM MgCl2, 2 mM dNTPs and BSA (Solis BioDyne, Estonia), 0.2 μM final concentration of each primer and 1 μl genomic DNA. The PCR profile consisted of an initial denaturation at 95 °C for 15 min followed by 25 cycles of denaturation at 94 °C for 60 s, annealing at 53 °C (set D) or 50 °C (set E) for 60 s, and extension at 72 °C for 60 s. Final extension of amplified DNA occurred at 72 °C for 10 min. All PCR reactions were carried out in a Mastercycler® ep Gradient thermal cycler (Eppendorf, Hamburg, Germany). Amplification fragments were separated on an ABI Prism 3130xl® Genetic Analyzer (Applied Biosystems, Foster City, CA, USA) and visualized with the softeare GeneMapper® ver. 3.5 (Applied Biosystems).
The nuclear SSR genotypes were encoded as binary data to facilitate analysis of the polyploid species S. alba and S. fragilis. Two of the utilised markers (SB38 and SB194) have been reported to have disomic-monogenic inheritance in tetraploid S. alba and S. fragilis ([16]), however this has not been demonstrated for the other SSR markers utilised in this study. The chloroplast marker data was combined into haplotypes. The pairwise Nei’s genetic distances between populations and varieties were calculated from the nuclear SSR data (encoded as binary data), and used to construct a neighbour-joining dendrogram. Identification of vegetatively propagated clones was done by simple matching of nuclear SSR fragments. Principal coordinates analysis (PCoA) was done based on the pairwise Nei’s genetic distance matrix. Genetic analyses and Mantel’s tests were done using the software package GenAlEx ver. 6.5 ([29]) and DARwin ([31]). The robustness of the dendrograms was examined by bootstrapping analysis (1000 bootstraps). Polymorphism information content (PIC) values were calculated as PIC = 1 - (p2-q2), where p is the fragment frequency and q is the no-fragment frequency ([26]). Dendrograms were constructed using the software MEGA 5.2 ([36]) and FigTree v 1.4.2 (⇒ http://tree.bio.ed. ac.uk/software/figtree/).
Results
Of the six analysed chloroplast markers, only two were polymorphic, which detected two polymorphic alleles each (CCMP3: 113bp and 114bp; CCMP10: 110bp and 111bp). These were combined into haplotypes, and only two haplotypes were identified within the analysed samples (designated as HapA and HapB). HapA was the most commonly identified haplotype, with 236 individuals (71%), while HapB was found in 95 individuals (29%). Most of the analysed stands contained individuals of only one haplotype, with the exception of the Vireši stand (HapA frequency 0.81) and the Vecmemele stand (HapA frequency 0.96). HapB was found in all individuals collected from S. fragilis and hybrid stands (Dviete, Piedruja, Lielindrica - S. fragilis, Ilukste). In addition, two individuals with HapB were identified from samples collected in the National Botanic Garden (’Vitellina’, S. × rubens).
Three of the analysed SSR markers amplified a maximum of two alleles per individual (SB38, SB194 and SB201), two of which (SB38 and SB194) were previously reported to have disomic-monogenic inheritance in tetraploid S. alba and S. fragilis ([16]). The remaining markers amplified more than two alleles in some of the analysed individuals, therefore the genotypes were analysed as dominant binary markers. The eight SSR markers identified a total of 75 fragments. Of these, 33 (44%) had a minor allele frequency of less than 0.05. The number of fragments detected in each population ranged from 25 (Ventspils) to 54 (Nica). The average number of fragments detected over all the analysed populations was 34.5. The average PIC values for the loci within each population ranged from 0.088 (Ilukste) to 0.231 (Nica). One unique allele was found in each of five populations (Laudona, Bebrene, Lielindrica - S. alba, Salaca, Kaunas), and two unique alleles were identified in the Nica population (Tab. 2). The Ventspils population was the least genetically diverse, and 14 of the 26 analysed individuals belonged to just three distinct clones. Evidence of vegetative propagation to a lesser extent was also found in Vireši (2 individuals, 1 clone), Ilukste (5 individuals, 2 clones) and Dviete (4 individuals, 2 clones).
Tab. 2 - Parameters of genetic diversity for the analysed populations.
| Population | N | No. of fragments |
No. of fragments (freq.≥ 5%) |
Mean PIC | No. of unique fragments |
|---|---|---|---|---|---|
| Nica (S.alba) | 24 | 54 | 46 | 0.231 | 2 |
| Ventspils (S.alba) | 26 | 25 | 21 | 0.093 | 0 |
| Vecmemele (S.alba) | 23 | 40 | 31 | 0.146 | 0 |
| Voleri/Riga (S.alba) | 24 | 38 | 35 | 0.163 | 0 |
| Laudona (S.alba) | 24 | 37 | 29 | 0.127 | 1 |
| Dviete (S.alba × S.fragilis) | 15 | 28 | 28 | 0.104 | 0 |
| Bebrene (S.alba) | 7 | 27 | 27 | 0.114 | 1 |
| Ilukste (S.fragilis) | 24 | 30 | 20 | 0.089 | 0 |
| Lielindrica (S.alba) | 24 | 35 | 26 | 0.115 | 1 |
| Lielindrica (S.fragilis) | 24 | 29 | 28 | 0.130 | 0 |
| Piedruja (S.fragilis) | 24 | 27 | 22 | 0.095 | 0 |
| Adaži (S.alba) | 18 | 38 | 38 | 0.156 | 0 |
| Vireši (S.alba) | 27 | 35 | 32 | 0.135 | 0 |
| Salaca (S.alba) | 21 | 33 | 24 | 0.124 | 1 |
| Kaunas (S.alba) | 17 | 42 | 42 | 0.178 | 1 |
| Mean | 21.786 | 34.533 | 29.071 | 0.130 | - |
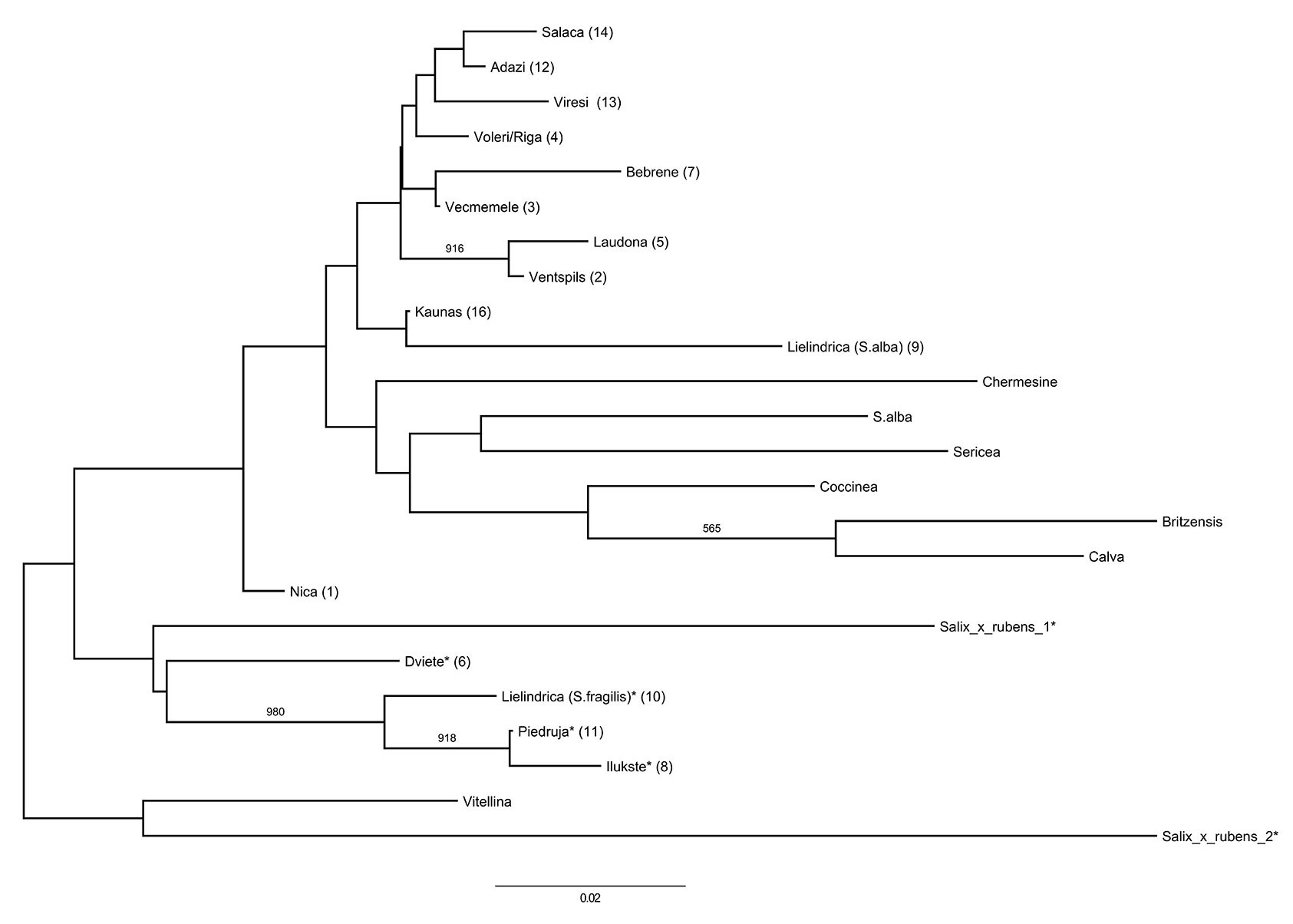
The calculated Nei’s genetic distance based on nuclear SSR marker genotypes, ranged from 0.006 (Piedruja-Ilukste) to 0.113 (Bebrene-Ilukste). The average pairwise Nei’s genetic distance was 0.051. Dendrogram analysis grouped the S. fragilis and hybrid populations into one cluster, and the S. alba populations into another (Fig. 4). However, only a few clusters had over 50% bootstrap support, and there was also no evidence of isolation by distance, with a Mantel test indicating no significant correlation between genetic distance and geographic distance, both for all populations and for the S. alba populations only. AMOVA results revealed some differentiation between the analysed populations, with 37% of the total genetic variation found between all populations (PhiPT = 0.371, p<0.001, 999 permutations). The separate analysis of S. alba populations revealed that between-population variability accounted for 25% of the total genetic variation (PhiPT=0.254, p<0.001, 999 permutations) The populations in the S. alba cluster show some geographical grouping, with the Salaca and Vireši populations found in northern Vidzeme, the Adaži and Voleri populations in the vicinity of Riga, and the Bebrene and Vecmemele populations found in the eastern region of Latvia (Fig. 3). One exception was the close genetic relationship of the Ventspils and Laudona population. These populations are located over 300 km distant from each other, but were among the most closely related pair of populations analysed, with a pairwise Nei’s genetic distance of 0.010. The Nica population was at an intermediate position between the two clusters.
Fig. 4 - Neighbour-joining dendrogram of Nei’s genetic distances between populations and varieties. (*): S. fragilis populations and hybrid varieties/individuals; (NBD): samples collected from the National Botanic garden. Numbers in brackets refer to populations labels reported in Tab. 1. Numbers on dendrogram branches indicate those having more than 50% bootstrap support (out of 1000 replications).
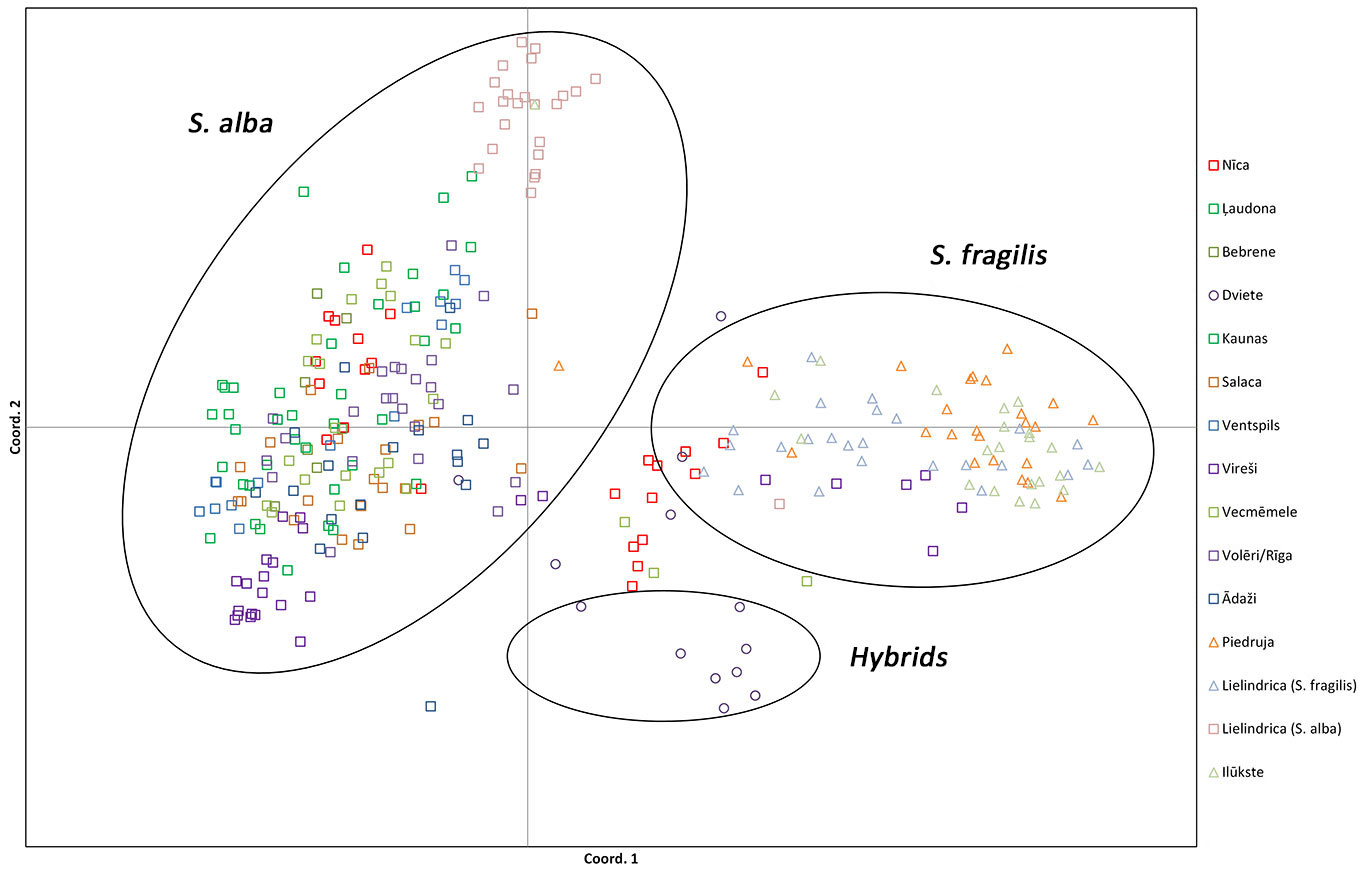
Principal coordinate analysis (PCoA) using the Nei’s genetic distance matrix calculated from nuclear marker data showed that the analysed individuals from the S. fragilis populations and putative hybrid individuals were mostly distinguished from the S. alba populations (Fig. 5). The putative hybrid individuals were intermediate between the two pure species clusters, but were closer to S. fragilis, corresponding to the neighbour-joining dendrogram. Twelve individuals from the Nica population were clustered with the S. alba individuals, while 12 individuals were clustered with the other S. fragilis/hybrid individuals according to the nuclear SSR marker data (Fig. 5), which is also reflected in the intermediate position of the Nica population in the neighbour-joining dendrogram. In addition, six individuals from the Vireši population, one from the Lielindrica (S. alba) and three individuals from the Vecmemele population were clustered with the S. fragilis/hybrid individuals. Of these 21 individuals (morphologically identified as S. alba, though clustering with S. fragilis/hybrid individuals based on nuclear SSR data), six had the chloroplast haplotype B, characteristic of S. fragilis (5 individuals from Vireši and one from Vecmemele).
Fig. 5 - Principal Coordinates Analysis of nuclear SSR genotype data. Individuals are labelled according to the population phenotype listed in Tab. 1 (S. alba, S. fragilis or hybrids).
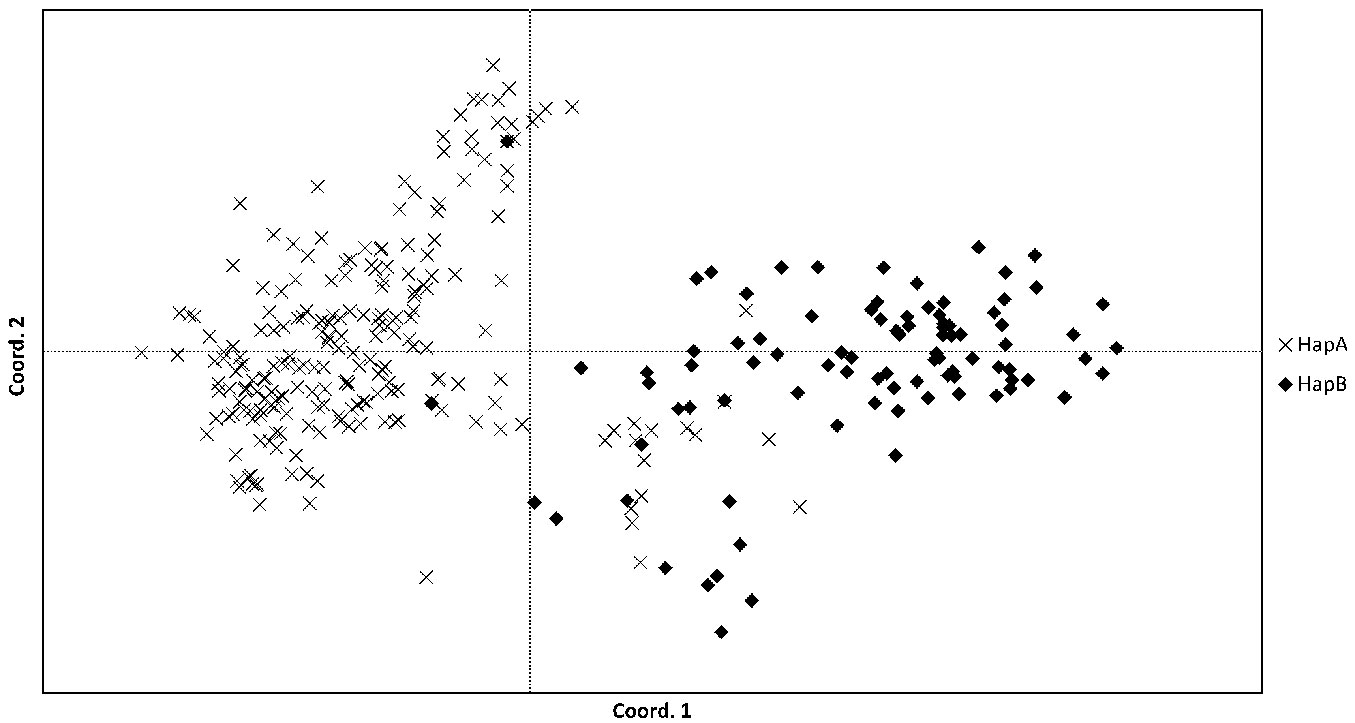
The individuals labelled as either HapA or HapB, as determined by the chloroplast markers, were clustered by PCoA using the nuclear SSR marker data (Fig. 6). The majority of the individuals distinctly cluster into two groups, reflecting their designation as S. alba or S. fragilis/hybrids. The majority of the hybrid individuals (identified phenotypically or by nuclear SSR marker data) as well as the S. fragilis populations were designated as HapB. However, there were 16 individuals which were grouped with the S. fragilis/hybrids individual that were designated as HapA. The majority of these (12 of the 16 individuals) were from the Nica population, three were from Vecmemele, one from Lielindrica (S. alba) population, and one sample from the botanic garden (S. × rubens). In addition, two individuals designated as HapB were clearly grouped with the S. alba individuals according to the nuclear SSR marker data (one individual from each of the populations Ilukste and Piedruja).
Fig. 6 - Principal Coordinates Analysis of nuclear SSR genotype data. Individuals are labelled according to chloroplast haplotypes HapA (predominantly S. alba) or HapB (predominantly S. fragilis/hybrids).
The individuals collected from the National Botanic Garden were split between the S. alba and S. fragilis/hybrid clusters. Most samples were clustered with the S. alba populations, however, the two individuals identified as Salix × rubens Shrank (S. alba × S. fragilis) clustered with the S. fragilis/hybrids, as well as the variety ’Vitellina’.
Discussion
The nuclear SSR data did not identify any highly differentiated S. alba populations in Latvia. The levels of diversity (number of fragments identified and average PIC) within each population were similar, even in the two populations with a smaller number of sampled individuals (Bebrene: 7 individuals; and Ventspils: 26 individuals), but with 14 analysed individuals belonging to just three clones, i.e., 15 genetically distinct individuals. This indicates a high level of genetic diversity within and between individuals in a single population, as expected from a dioecious species with wind dispersed seeds ([37]).
In previous studies of S. alba and S. fragilis, a higher differentiation of these two species was reported by using DNA markers as compared to morphological traits ([38]). It has been suggested that this might be a consequence of the allotetraploid nature of the genomes of S. alba and S. fragilis, with the two species sharing one common or two highly related diploid ancestor species, but differentiated from each other by the genomic contributions of divergent second diploid ancestor species ([2]). The allotetraploid genomes of both species, may suggest disomic segregation of genetic loci. However, the SSR markers utilised in this study were developed from Salix burjatica, and have been shown to amplify in a wide range of other Salix species ([3]), thus explaining why more than two alleles were amplified at some of the analysed SSR loci.
Only two chloroplast haplotypes were identified within the analysed individuals, and the same two haplotypes were also identified in the cultivated clones and varieties collected from the National Botanic Garden, which are presumably of different provenance (not from the Baltic region). Analysis with additional chloroplast markers, which reveal further polymorphism/haplotypes, may allow a more fine-scale elucidation of the provenance of the Baltic S. alba populations. Analysis of additional reference material sourced from non-Baltic countries will allow a better understanding of the extent of chloroplast polymorphism revealed by the utilised markers and the levels of diversity within S. alba and S. fragilis at the European scale. The chloroplast haplotypes were able to differentiate the majority of S. alba (HapA) and S. fragilis (HapB) individuals. The majority of putative hybrid individuals had the S. fragilis haplotype (HapB), suggesting that the in general hybridisation events occur by fertilisation of S. fragilis mothers by S. alba pollen; however, there may be a lower frequency of reciprocal fertilisation of S. alba by S. fragilis pollen. Alternatively, there may be a low frequency of shared chloroplast haplotypes between S. alba and S. fragilis due to natural hybridisation events. A mixture of haplotypes was found only in two populations, Vireši and Vecmemele. The majority of individuals in these populations with the S. fragilis haplotype (HapB) also clustered with the S. fragilis/putative hybrid individuals according to the nuclear SSR marker data, while the individuals from these populations that clustered with the S. alba individuals based on nuclear SSR data also had the S. alba chloroplast haplotype (HapA). This suggests that hybridisation can occur in naturally established and regenerated stands or alternatively, that S. alba and S. fragilis individuals are growing in mixed stands, and that the morphological parameters that distinguish these species are homogenised by environmental and other growing site characteristics. However, the Nica population was homogeneous for the S. alba chloroplast haplotype (HapA), but was split into two clusters according to the nuclear SSR data. This suggests that this population may contain some individuals generated by hybridisation in the opposite direction (i.e., fertilisation of S. alba mother trees by S. fragilis pollen), as compared to the majority of the hybrids identified in this study (fertilisation of S. fragilis mother trees by S. alba pollen). This population is located in the south-western region of Latvia, which has a relatively higher density of S. fragilis and also has a less continental climate than eastern Latvian regions. Such factors may influence the hybridisation pattern, including effects on the phenology of these species, for instance alteration of flowering times. These putative hybridisation occurrences should be analysed in more detail at a molecular and phenotypic level by analysing additional putative hybrid individuals, particularly in stands where a mixture of chloroplast haplotypes and morphological characteristics are found.
In general, the nuclear SSR genotypic data, the chloroplast haplotypes and the phenotypic assessments of the analysed populations were well correlated. The individuals that were phenotypically identified as putative hybrids were clustered between the S. alba and S. fragilis individuals using the nuclear SSR markers, while the majority of these individuals had the chloroplast haplotype B (HapB) predominant in S. fragilis. The individuals sampled from the National Botanic Garden were genetically similar to the S. alba populations, with the exception of the individuals previously identified as Salix × rubens Shrank (hybrid of S. alba and S. fragilis) as well as the cultivar ’Vitellina’. The two Salix × rubens Shrank individuals were designated as having different haplotypes (HapA and HapB), indicating that the hybridisation direction may differ for each individual. Alternatively, the haplotype identified primarily in S. fragilis and hybrid individuals (HapB), also found at a lower frequency in S. alba populations, could be the result of natural hybridisation, as reported previously for other Salix species ([28]). Previous investigations of the willow complex S. alba - S. × rubens - S. fragilis using DNA markers ([7]) revealed that S. × rubens can be separated into two groups (S. × rubens and S. × rubens var. basfordiana), which clustered with S. alba and S. fragilis, respectively. A more precise taxonomic classification for the two S. × rubens individuals analysed in this study was not available, but the alternating chloroplast haplotypes in these two individuals may also support the results reported by De Cock et al. ([7]). The individual identified as ’Vitellina’ which was collected from the National Botanic Garden, clustered with the S. fragilis and hybrid individuals according to the nuclear SSR data and this individual also had the chloroplast haplotype predominantly found in S. fragilis and hybrid individuals (HapB). This was surprising, as ’Britzensis’ is reported as a cultivar of the S. alba sub-species vitellina or variety ’Vitellina’. However, the ’Britzensis’ individual analysed was genetically similar to the S. alba individuals, and also had the S. alba chloroplast haplotype (HapA), in contrast to the ’Vitellina’ individual analysed in this study. A more detailed morphological assessment of the sampled ’Vitellina’ individual and further genetic analysis of additional ’Vitellina’ individuals could help determine if this particular individual was mislabelled, or if other representatives of the variety ’Vitellina’ also are genetically distinct from S. alba.
This is the first study which examined the population structure of S. alba and its hybrids in Latvia. Analysis of the S. alba individuals collected from the National Botanic Garden, which represent a range of cultivated forms and varieties, as well as hybrids, indicated that the individuals designated as S. alba sub-species, were genetically similar to the analysed S. alba natural populations, and that there is little genetic differentiation between the S. alba populations and sub-species, supporting the hypothesis that the classification of cultivated clones or decorative forms as sub-species and varieties is not taxonomically significant. Further analysis of natural S. alba populations from beyond the Baltic region will indicate if the level of genetic differentiation in S. alba is low at the European scale, as found for other broadleaf tree species, e.g., Fraxinus excelsior L. ([12]). The analysis of additional nuclear and chloroplast markers may also allow better differentiation of S. alba populations.
Webb ([40]) proposed a set of criteria for determining the native status of a species: fossil evidence, historical evidence, habitat, geographical distribution, frequency of known naturalization, genetic diversity, reproductive pattern, possible means of introduction. Consideration of these criteria suggests that the majority of the examined S. alba populations are probably natural. There are no fossil records available of this species from Latvia, but historical records report that this species has been described as growing in natural stands in the territory of Latvia for over 200 years ([10], [17]). The habitats where the majority of the analysed S. alba populations were collected were assessed as natural (with the exception of the Ventspils population), and this species is distributed throughout the territory of Latvia ([20] - Fig. 2). The genetic diversity analyses did not identify any highly differentiated populations; however, the discriminative power of the used nuclear and chloroplast markers may not be sufficient to detect possible introductions, as the presumably foreign clones in the National Botanic Garden collection were not differentiated from the analysed populations. In this regard, our results are not conclusive, though it seems unlikely that all the analysed populations are introduced, particularly as Latvia is already on the border of the previously described distribution area for this species. There was only a small number of genetically identical individuals identified, indicating a low level of vegetative reproduction (again with the exception of the Ventspils population). Although there is a long history of cultivation and planting of S. alba in parks and gardens in Latvia ([4]), which would provide a plausible means of introduction, the majority of the criteria suggest a natural origin of S. alba in Latvia. This supports the hypothesis that Latvia is within the natural distribution range of S. alba, thus many of the natural stands found in Latvia could be considered to be autochthonous.
Our results provide the basis for further, more detailed investigations of S. alba populations in Latvia, allowing the identification of possibly introduced or artificially regenerated stands, and the determination of potential S. alba genetic resource stands. Within this study, the Ventspils population, which was suspected to be at least partially artificially established from in situ observations, was confirmed to contain a high proportion of vegetatively propagated individuals, which could be indicative of reproduction via cuttings. In addition, these results confirm that S. alba hybridises with S. fragilis, and that hybrid individuals can be found in natural stands. The results of the chloroplast marker analysis indicate that the predominant hybridisation occurs by fertilisation of S. alba mother trees by S. fragilis pollen, however, the extent of haplotype sharing between these two species should be further investigated. In addition, stands with varying proportions of chloroplast haplotypes were identified in this study, and these stands should be examined in more detail using both molecular and phenotypic methods.
Acknowledgements
This research was funded within the ERDF project no. 2014/0025/2DP/2.1.1.1.0/ 14APIA/VIAA/101 “Development of identification technologies for vegetatively propagated short rotation tree species”
References
CrossRef | Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Authors’ Info
Authors’ Affiliation
Maris Laivins
Agnese Gailite
Anna Korica
Dagnija Lazdina
Vilnis Skipars
Ilze Veinberga
Latvian State Forest Research Institute “Silava”, 111 Rigas st, Salaspils, LV-2169 (Latvia)
Corresponding author
Paper Info
Citation
Rungis D, Laivins M, Gailite A, Korica A, Lazdina D, Skipars V, Veinberga I (2017). Genetic analysis of Latvian Salix alba L. and hybrid populations using nuclear and chloroplast DNA markers. iForest 10: 422-429. - doi: 10.3832/ifor2004-009
Academic Editor
Fikret Isik
Paper history
Received: Feb 05, 2016
Accepted: Nov 27, 2016
First online: Mar 24, 2017
Publication Date: Apr 30, 2017
Publication Time: 3.90 months
Copyright Information
© SISEF - The Italian Society of Silviculture and Forest Ecology 2017
Open Access
This article is distributed under the terms of the Creative Commons Attribution-Non Commercial 4.0 International (https://creativecommons.org/licenses/by-nc/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Web Metrics
Breakdown by View Type
Article Usage
Total Article Views: 50738
(from publication date up to now)
Breakdown by View Type
HTML Page Views: 41995
Abstract Page Views: 3529
PDF Downloads: 3984
Citation/Reference Downloads: 23
XML Downloads: 1207
Web Metrics
Days since publication: 3325
Overall contacts: 50738
Avg. contacts per week: 106.82
Article Citations
Article citations are based on data periodically collected from the Clarivate Web of Science web site
(last update: Mar 2025)
Total number of cites (since 2017): 4
Average cites per year: 0.44
Publication Metrics
by Dimensions ©
Articles citing this article
List of the papers citing this article based on CrossRef Cited-by.
Related Contents
iForest Similar Articles
Technical Reports
Conservation and use of elm genetic resources in France: results and perspectives
vol. 13, pp. 41-47 (online: 03 February 2020)
Technical Reports
Population genetic structure of Platanus orientalis L. in Bulgaria
vol. 4, pp. 186-189 (online: 11 August 2011)
Research Articles
Delineation of seed collection zones based on environmental and genetic characteristics for Quercus suber L. in Sardinia, Italy
vol. 11, pp. 651-659 (online: 04 October 2018)
Editorials
Workshop COST E52 “Evaluation of beech genetic resources for sustainable forestry”
vol. 2, pp. 104 (online: 10 June 2009)
Research Articles
Growth, spring phenology and stem quality of four broadleaved species assessed in provenance trials in the Netherlands - Implications for seed sourcing
vol. 18, pp. 242-251 (online: 22 September 2025)
Research Articles
Networking sampling of Araucaria araucana (Mol.) K. Koch in Chile and the bordering zone of Argentina: implications for the genetic resources and the sustainable management
vol. 2, pp. 207-212 (online: 22 December 2009)
Research Articles
Fine-scale spatial genetic structure in a multi-oak-species (Quercus spp.) forest
vol. 8, pp. 324-332 (online: 05 September 2014)
Commentaries & Perspectives
The genetic consequences of habitat fragmentation: the case of forests
vol. 2, pp. 75-76 (online: 10 June 2009)
Research Articles
Some refinements on species distribution models using tree-level National Forest Inventories for supporting forest management and marginal forest population detection
vol. 11, pp. 291-299 (online: 13 April 2018)
Research Articles
Patterns of genetic diversity in European beech (Fagus sylvatica L.) at the eastern margins of its distribution range
vol. 10, pp. 916-922 (online: 10 December 2017)
iForest Database Search
Search By Author
Search By Keyword
Google Scholar Search
Citing Articles
Search By Author
Search By Keywords
PubMed Search
Search By Author
Search By Keyword