Estimation of aboveground forest biomass in Galicia (NW Spain) by the combined use of LiDAR, LANDSAT ETM+ and National Forest Inventory data
iForest - Biogeosciences and Forestry, Volume 10, Issue 3, Pages 590-596 (2017)
doi: https://doi.org/10.3832/ifor1989-010
Published: May 15, 2017 - Copyright © 2017 SISEF
Research Articles
Abstract
Assessing biomass is critical for accounting bioenergy potentials and monitoring forest ecosystem responses to global change and disturbances. Remote sensing, especially Light Detection and Ranging (LiDAR) data combined with field data, is being increasingly used for forest inventory purposes. We evaluated the feasibility of the combined use of freely available data, both remote sensing (LiDAR data provided by the Spanish National Plan for Aerial Ortophotography - PNOA - and Landsat vegetation spectral indices) and field data (from the National Forest Inventory) to estimate stand dendrometric and aboveground biomass variables of the most productive tree species in a pilot area in Galicia (northwestern Spain). The results suggest that the models can accurately predict dendrometric and biomass variables at plot level with an R2 ranging from 0.49 to 0.65 for basal area, from 0.65 to 0.95 for dominant height, from 0.48 to 0.68 for crown biomass and from 0.55 to 0.82 for stem biomass. Our results support the use of this approach to reduce the cost of forest inventories and provide a useful tool for stakeholders to map forest stand variables and biomass stocks.
Keywords
Biomass Maps, Forest Inventory, LiDAR, Landsat Vegetation Indices
Introduction
Measurement and mapping of aboveground forest biomass at diverse scales is critical for estimating global carbon storage and assessing those ecosystem responses to climate change and anthropogenic disturbance ([38]). Estimation of forest biomass is of great importance because of the increasing value of this renewable resource for energy production ([15]). Furthermore, biomass estimation and mapping are used to quantify fuel loads for fire risk assessment and fire prevention planning purposes ([16]).
The method most commonly used to estimate aboveground forest biomass is the forest inventory based on plot data. However, given the high costs and operational difficulties associated with this method, the use of remotely sensed data in combination with field-work is becoming increasingly popular for forest inventory purposes ([19]). Moreover, remote-sensing methods enable the estimation of stand dendrometric and biomass values at each pixel location, rather than estimation of average or total biomass within a given area ([18]).
Most studies using remote-sensing methods have been based on medium-to-high resolution sensors, especially the Landsat-TM sensor, given the good compromise between spectral and temporal resolution of the data achieved ([19], [29]). Remote-sensing indices have been used as explanatory variables for models of forest biomass and productivity ([30]). LiDAR (Light Detection and Ranging) technology has been recognized as a much more efficient, accurate and cost effective approach to sensing aboveground biomass. This technology is generally regarded as a more accurate method because LiDAR sensors provide information about vertical height of individual pulses returns, which can then be used to predict canopy attributes ([10]). Previous studies have demonstrated the success of LiDAR estimates of aboveground biomass based on the relationship between LiDAR metrics and field measurements of biomass obtained from allometric models ([45]). In this study we estimated aboveground forest biomass in Galicia (NW Spain) by the combined use of LiDAR, LANDSAT ETM+ and field data.
Two main types of statistical approaches are applied to estimate biomass from remotely sensed data: one focused on the data and the other on algorithms ([6]). The first assumes a stochastic model (linear, nonlinear regressions) used both to predict responses of population units not in the sample and future responses ([19], [14], [15]). The second approach (generally called machine-learning models) has been reported for implicitly inferring unknown relationships underlying a given dataset, being versatile enough to uncover complicated non-linear relationships ([11]). In the present study the first approach was chosen.
Forest inventories based on remotely-sensed data can be carried out at two different spatial scales: the individual tree level (associated with spatially dense LiDAR data, > 5 points m-2 - [22]) and the area-based, which can also be used with low density LiDAR data ([45], [14], [15]). In this study, we focused on area-based forest inventories, as we used low density LiDAR data provided by the Spanish National Plan for Aerial Ortophotography (PNOA) for the Spanish territory (0.5 pulse m-2), being higher point densities required for the estimation at individual tree level ([43]). The use of LiDAR for aboveground biomass estimations is still considered a challenging task, especially when using low density scanning systems, due to its lower accuracy in comprehensively reflecting the canopy structure ([27]). However, low density LiDAR has been shown to be quite effective for predicting biomass and other variables in temperate forests ([36], [39], [33]).
For large areas covered by different types of vegetation, models based on remote sensing data are usually developed for each vegetation type, as this kind of models is greatly influenced by species and stand structure ([36], [39], [24]). Implementation of models for mapping stand dendrometric and biomass variables requires an accurate classification of the area ([36], [39], [24]).
Galicia has one of the greatest potential forest production rates in Europe, varying from 6 to 12 m3 ha-1 year-1 depending on the particular species ([8]) and producing 45% of Spain’s timber and 4.5% of Europe’s. The most productive and fast-growing tree species in Galicia are Eucalyptus spp. (Eucalytus globulus Labill. and Eucalyptus nitens Deane et Maiden), Pinus pinaster Ait. and Pinus radiata D. Don, all of which are grown in extensive commercial forest plantations to produce panelboard, sawlog and pulpwood ([34]). Stands dominated by these tree species account for 133.224.356 m3 of standing timber (69.1% of the total tree standing timber volume) in Galicia, and cover an area of 896.342 ha (62.9% of the total tree-covered area) in the region ([31]).
The objective of this study was to evaluate the feasibility of the combined use of freely available remote sensing (low density LiDAR and Landsat) and National Forest Inventory data combined with the National Forest Map (based on PNOA aerial ortophoto, with a scale of 1:25.000 and minimum mapping unit of 1 ha) to estimate stand dendrometric and aboveground biomass variables of the most productive tree species in a pilot area in Galicia. This is the first study using this freely available information for these tree species in the NW Spain. The results may also be applicable to similar areas covered by similar sensors.
Material and methods
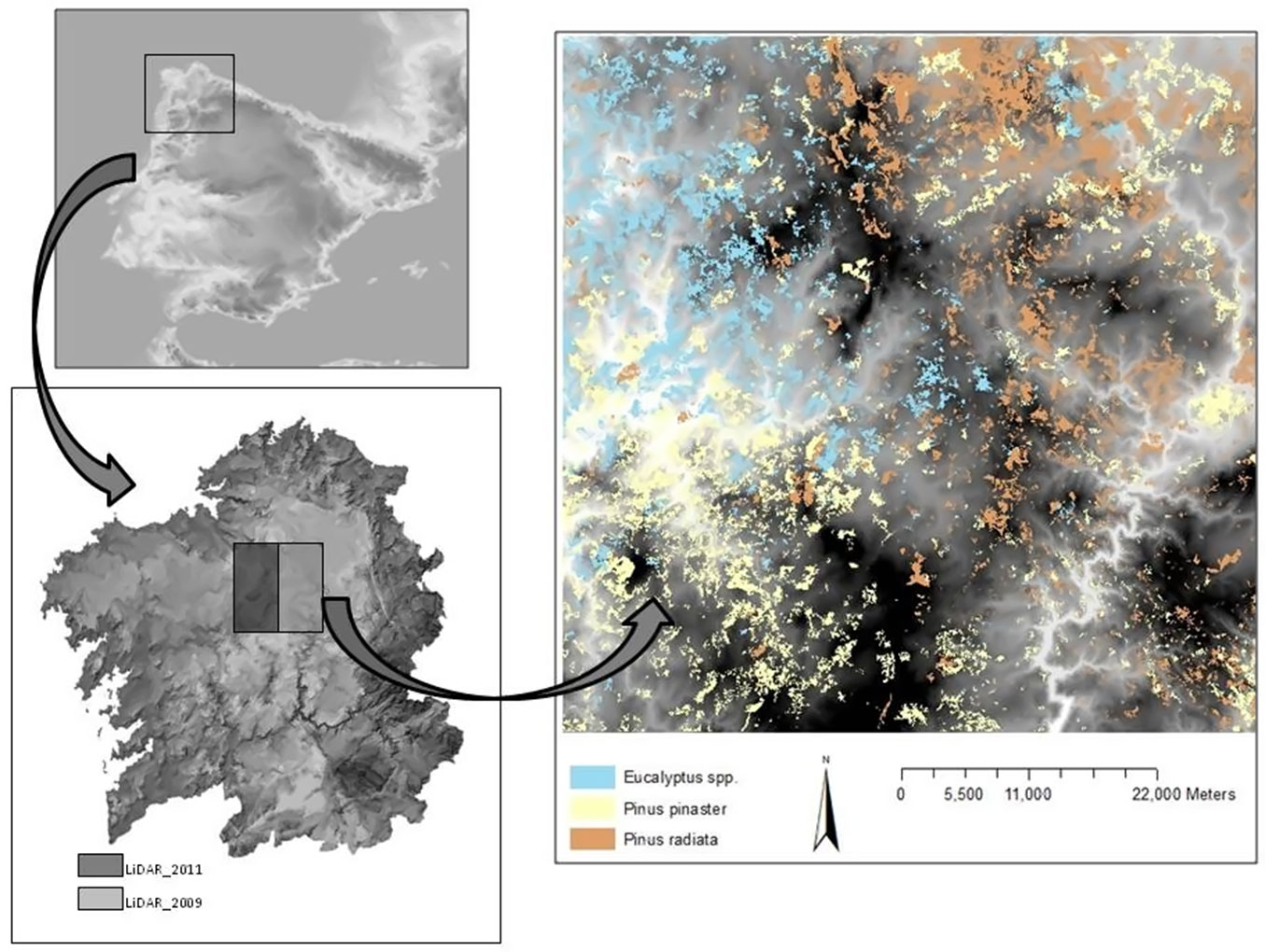
The study was conducted in the municipality of Palas de Rei (Lugo), in northwestern Spain (42° 52′ 23″ N; 07° 52′ 08″ W - Fig. 1). The study area consisted of a 60 × 60 km square, centered in Palas de Rei encompassing parts of the provinces of Lugo, Pontevedra and A Coruña. The mean elevation of the area is 525 m (range 147-1179 m) and the mean slope is 11.3%. The climate is Mediterranean, with a continental influence. The average precipitation is 1188 mm year-1, and the mean annual temperature is 12.3 °C, with maximum temperatures in August (19.0 °C) and minimum temperatures in January (6.8 °C). Forests (more than 20% of tree cover) occupy 150.396.6 ha of the total surface area. The main tree species in the area are Pinus pinaster (covering 27.8% of the total forest area), Pinus radiata (22.7%), Quercus robur (30.7%) and Eucalyptus spp. (mainly E. nitens and E. globulus - 13.8%).
Fig. 1 - Location of the study area and National Forest Map polygons dominated by Pinus pinaster, Pinus radiata and Eucalyptus spp.
Field data
The Fourth Spanish National Forest Inventory (SNFI-4) plots were used as the source of field data for this study. Plots within the study area dominated by the three selected tree species, with tree cover > 20% and presence of trees with diameter at breast height (dbh) > 7.5 cm, were selected for the study ([31]). The plots were established at the intersections of a 1 × 1 km grid, totaling 873 plots over the whole study area. In 749 of these, tree cover was > 20%, with presence of trees with dbh > 7.5 cm. In total, 159 plots were dominated by Pinus pinaster, 142 by Pinus radiata and 109 by Eucalyptus spp. The measurements were carried out during 2008 and 2009 (Tab. 1).
Tab. 1 - Mean (± standard error) tree density, basal area (G), dominant height (H0), crown biomass (Wcr), stem biomass (Wst) per sample plot of the National Forest Inventory within the study area for each species.
| Main species |
Density (trees ha-1) |
G (m2 ha-1) |
H0 (m) |
Wcr (Mg ha-1) |
Wst (Mg ha-1) |
|---|---|---|---|---|---|
| Eucalyptus spp. (n=109) | 783 ± 48 | 17.0 ± 1.4 | 25.5 ± 1.1 | 17.1 ± 1.6 | 89.9 ± 10.0 |
| Pinus pinaster (n=159) | 579 ± 39 | 17.3 ± 1.1 | 20.6 ± 0.6 | 19.6 ± 1.2 | 58.4 ± 4.1 |
| Pinus radiata (n=142) | 646 ± 36 | 19.9 ± 1.1 | 20.4 ± 0.6 | 20.0 ± 1.1 | 70.8 ± 4.8 |
Sample plots consisted of four concentric circles of radii 5, 10, 15 and 25 m, in which dbh and total height were measured in all trees of dbh > 7.5, 12.5, 22.5 and 42.5 cm, respectively. Diameter at breast height was measured to the nearest 0.1 cm with a graduated caliper, and tree height was measured to the nearest 0.1 m with a hypsometer. The number of stems per hectare, basal area and dominant height (mean height of the 100 thickest trees per hectare) were calculated from the tree variable measurements by the use of expansion factors. These factors express the number of trees per hectare that each measured tree represents in the inventory in relation to the subplot radius.
Aboveground biomass per plot was estimated by applying existing allometric models for each measured tree within the plot. Specific allometric models were used for each species, and models constructed for the same ecoregion (NW Spain) were selected. The explanatory variables included in the models were tree variables measured in the National Forest Inventory (dbh and tree height). The allometric models by the following authors were used for the different species: Jiménez et al. ([20]) and Gómez-Vázquez et al. ([13]), for Pinus pinaster; Balboa-Murias et al. ([2]), for Pinus radiata; Brañas et al. ([5]) for Eucalyptus globulus; and Pérez-Cruzado & Rodríguez-Soalleiro ([40]) for Eucalyptus nitens. For other tree species inside the plots, we used the model by Balboa-Murias ([1]) for Quercus robur and those reported by Montero et al. ([35]) for other species. After applying the models, we estimated the biomass values for following components in each tree measured: leaves, stem, fine branches (diameter < 2 cm) and coarse branches (diameter ≥ 2 cm). To obtain the aboveground biomass at plot level, we used the above-mentioned expansion factor. Aboveground biomass at plot level was subdivided in two components: crown biomass (leaves and branches) and stem biomass (Tab. 1).
The National Forest Map, associated with the National Forest Inventory was obtained from the PNOA aerial ortophoto (scale 1:25.000, minimum mapping unit of 1 ha) and used to classify vegetation types and spatially define the polygons dominated by each of the species under study (Fig. 1).
LiDAR data
The LiDAR data were provided by the PNOA. The study area was surveyed at two different times: once in 2009 (province of Lugo) and then in 2011 (provinces of Pontevedra and A Coruña). Data were delivered in 2 × 2 km tiles of points in LAS binary files. The resulting LiDAR point density for the study area was 0.5 pulse m-2, with a vertical accuracy greater than 0.2 m. A total of 961 LAS files were required in order to cover the study area.
The LiDAR data were processed using the software FUSION LDV 3.50 ([32]). After eliminating the noise from the point cloud, the Digital Elevation Model (DEM) of the study area was obtained. This was done by first filtering the ground returns (using the “GroundFilter” tool) by implementing a filtering algorithm, adapted from Kraus & Pfeiffer ([23]). The DEM (1 m spatial resolution) was then generated using these returns through the “GridSurfaceCreate” command and used to normalize the heights of the point cloud. The LiDAR height and intensity statistics for each sample plot were obtained using the “ClipData” and “CloudMetrics” commands and the plot boundaries (25 m radius). A predefined threshold of 2 m above the ground was applied in order to exclude returns not corresponding to crowns (e.g., understory, rocks, shrubs).
Landsat ETM+ data
We obtained a cloud-free Landsat ETM+ scene corresponding to a date close to the National Forest Inventory and LiDAR survey (1 June 2009). Digital numbers were converted to radiometric values by using the specific gain and offset of the sensor. Reflectance was obtained using the method of NASA ([37]). The images (SLC-off) were fused using the method described by Scaramuzza et al. ([42]). Reflectance values were used to obtain several vegetation indices (Tab. 2) for each plot sampled.
Tab. 2 - Landsat vegetation spectral indices employed in the study. (ρNIR): Near infrared; (ρR): Red; (ρB): Blue.
| Index | Equation | Reference |
|---|---|---|
| Simple Ratio (SR) | SR = ρ NIR / ρ R | Jordan ([21]) |
| Normalized Difference Vegetation Index (NDVI) | NDVI = (ρNIR - ρR)/ (ρNIR + ρR) | Rouse et al. ([41]) |
| Soil-Adjusted Vegetation Index (SAVI) | SAVI = (1+0.5) · (ρNIR - ρR)/ (ρNIR + ρR + 0.5) | Huete ([17]) |
| Normalized Ratio Vegetation Index (NRVI) | NRVI = (ρR/ρNIR - 1)/ (ρR/ρNIR + 1) | Baret & Guyot ([3]) |
| Enhanced Vegetation Index (EVI) | EVI = 2.5 · (ρNIR - ρR)/(ρNIR + 6 · ρR + 7.5 · ρB + 1) | Liu & Huete ([28]) |
Regression models
Linear, power function and exponential models were used to estimate how plot values (stand variables: basal area and dominant height; aboveground biomass: crown and stem biomass) were related to LiDAR variables (height and intensity metrics) and Landsat vegetation indices. Models were obtained for each dominant species (Pinus pinaster, Pinus radiata and Eucalyptus spp.) and for each LiDAR survey area (2009 and 2011). Separate models were constructed for different species because they have different structural characteristics (e.g., tree architecture, canopy stratification, canopy density, etc.) that generate differences in the models ([10], [11]), and because the accuracy of prediction depends on the type of forest ([10]). The LiDAR surveys were separated because the quality of laser-derived data depends on flight height, scan angle, point density and footprint size, among other factors. In the case of Eucalyptus spp., we only used the LiDAR_2011, as only a few of the National Forest Inventory plots dominated by this species were surveyed in 2009. The model expressions are as follows (eqn. 1, eqn. 2, eqn. 3):
where Y represents the field values (stand dendrometric and aboveground biomass variables), Xi represents a set of m independent variables (height and intensity LiDAR variables and Landsat vegetation indices), α and βi (i = 1, …, m) are parameters to be estimated, and ε is the error term.
Linear models were fitted by stepwise regression. Power and exponential models were fitted by nonlinear regression after prior linearization (taking natural logarithms) to select (by linear regression) the most significant subset of independent variables to include in the model. To correct for bias in log-transformed allometric equations, we adopted a correction factor ([44]). Heterocedasticity and multicollinearity among explanatory variables were checked. The presence of multicollinearity among variables was evaluated by the condition number ([4]), selecting the models with condition number smaller than 10 (collinearity is not a major problem). For each dominant tree species and LiDAR survey, the selected model included the combination of independent variables with the largest coefficient of determination (R2, defined as the square correlation coefficient between the measured and estimated values), the smallest values of the Akaike’s information criterion (AIC - [7]) and the root mean squared error of the estimate (RMSE).
Raster files were obtained for each explanatory variable (for LiDAR variables through the “CSV2Grid” FUSION command) with a spatial resolution similar to the Landsat image (30 m). The constructed models were used for spatial extrapolation of stand dendrometric and aboveground biomass variables to the study area, and the Spanish Forest Map was used to identify polygons dominated by the species considered. To avoid the inclusion of bias in the spatial extrapolation using model predictions, model-assisted estimators were employed to include a correction for estimated bias ([33]). This estimate is adjusted for deviations between the model predictions and the observed values in the sample (eqn. 4):
where MA mean is the model-assisted regression estimator of means, N is the population size, hat{y}i is obtained from the models using the model parameters estimates and ε = 0. The first term in the equation is the mean of the model predictions (hat{y}i) for all population units, and the second term is an estimate of bias calculated over the sample units and compensates for systematic model prediction errors.
Results
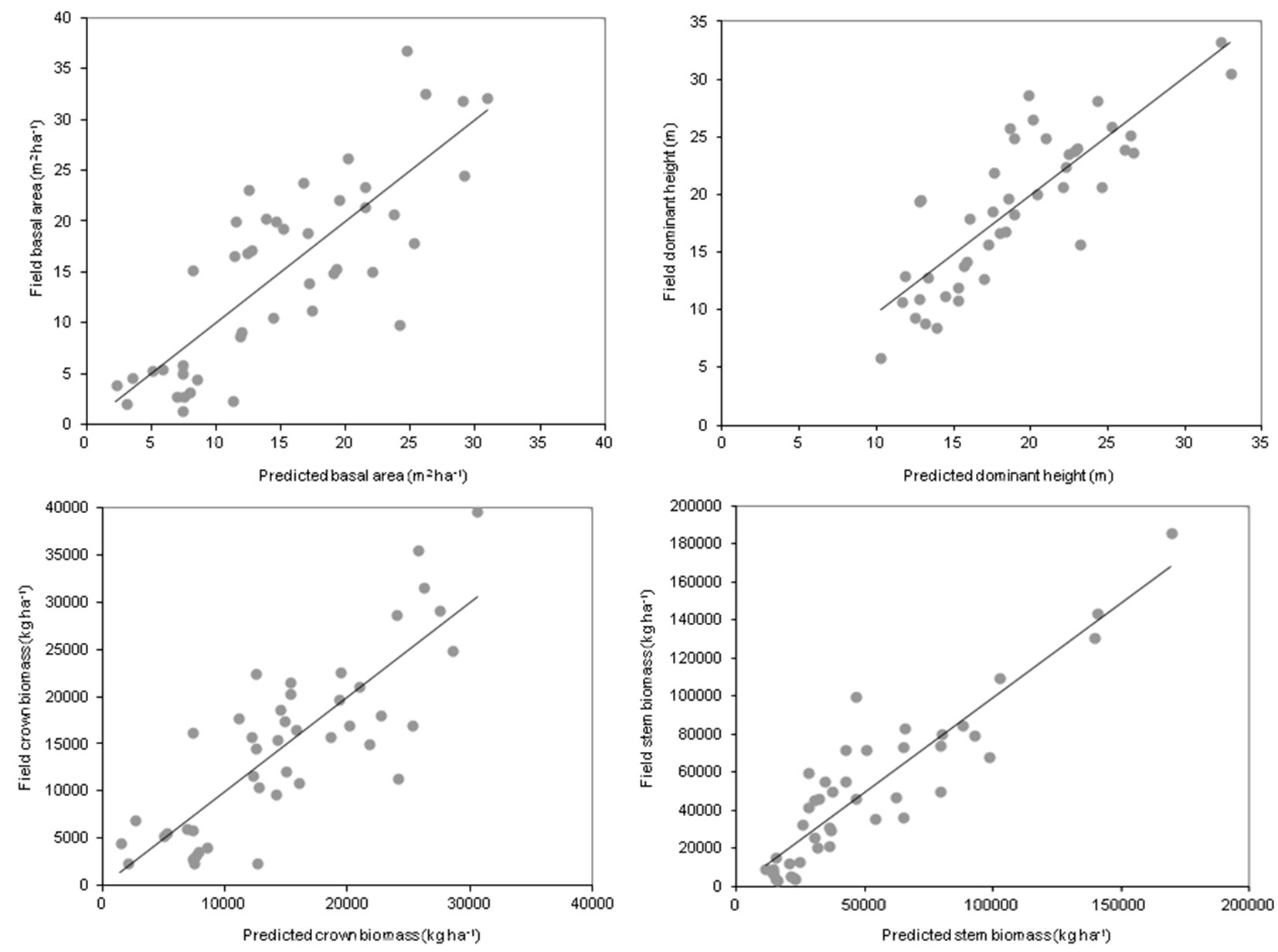
The models obtained for each dominant species and LiDAR survey, the goodness-of-fit-statistics for the most significant model constructed and the stand dendrometric and aboveground biomass variables are shown in Tab. 3. Linear and exponential models were selected on the basis of their performance. Eucalyptus spp. stands produced the least quality of fit in terms of R2, with models providing values of 49% for basal area, 65% for dominant height, 48% for crown biomass and 55% for stem biomass. By contrast, Pinus radiata stands (LiDAR_2011 data) produced the greatest quality of fit for basal area (65%), crown biomass (68%) and stem biomass (82%) (Fig. 2). The greatest quality of fit for dominant height was obtained for Pinus pinaster (LiDAR_2009 data), with a value of 95%.
Tab. 3 - Results of basal area (G), dominant height (H0), crown biomass (Wcr), stem biomass (Wst) modelling. (RMSE): root of mean squared error; (AIC): Akaike’s information criterion; (LIN): linear models; (EXP): exponential models; (Elmax): Maximum height; (1st Ret above 2): Number of first return above 2 m; (IntL3): L3 moments of intensity; (Int L4): L4 moments of intensity; (Elp90): Height 90th pertencile value; (Elp95): Height 95th pertencile value; (Elskewness): Height skewness; (Perc Ret above mode): Percentage of all returns above the mode height; (Ret above mean): number of returns above the mean value; (Ret above 2): number of returns above 2 m; (Elp10): height 10th percentile value; (ElCURTmenaCUBE): Cubic mean; (Ret2): Number of second returns; (IntLskewness): Moment ratio of intensity skewness; (ElCV): Coefficient of variation of height; (Ret3): Number of third returns; (Elp30): height 30th pertencile value; (Elp40): Height 40th pertencile value; (Intmode): Intensity mode; (Elp75): Height 75th pertencile value; (Elp01): height 1st pertencile value; (Int01): Intensity 1st pertencile value; (SR): Landsat Simple Ratio; (SAVI): Landsat SAVI Index; (ElL4): Moment L4 of height.
| Main species | Variable | Model | R2 | RMSE | AIC |
|---|---|---|---|---|---|
| Eucalyptus spp. LiDAR_2011 (n=85) |
G (m2 ha-1) | EXP (Elmax - 1st Ret above 2 - IntL3) | 0.49 | 8.7 | 334 |
| H0 (m) | EXP (Elmax - IntL4) | 0.65 | 5.9 | 273 | |
| Wcr (Mg ha-1) | EXP (Elmax) | 0.48 | 10.2 | 1390 | |
| Wst (Mg ha-1) | LIN ( Elmax - Elp90 - Elp95 - Elskewness) | 0.55 | 58.0 | 1657 | |
| Pinus pinaster LiDAR_2009 (n=54) |
G (m2 ha-1) | LIN (Elp95 - 1st Ret above 2) | 0.62 | 9.2 | 168 |
| H0 (m) | LIN (Elp95 -Perc Ret above mode - Elp90) | 0.95 | 1.3 | 27 | |
| Wcr (Mg ha-1) | LIN (Elp95 - Ret above mean) | 0.59 | 10.8 | 677 | |
| Wst (Mg ha-1) | EXP (Elp95 -Ret above 2 - Elp10) | 0.65 | 33.7 | 761 | |
| Pinus pinaster LiDAR_2011 (n=105) |
G (m2 ha-1) | EXP (ElCURTmeanCUBE - Ret2 - IntLskewness - ElCV) | 0.61 | 7.2 | 406 |
| H0 (m) | LIN (Elmax - Elskewness) | 0.73 | 3.9 | 283 | |
| Wcr (Mg ha-1) | EXP (Elmax - Ret3 - IntLskewness - Elp30 - IntL4) | 0.58 | 8.4 | 1822 | |
| Wst (Mg ha-1) | EXP (Elmax - Ret2 - IntLskewness - Elp40) | 0.60 | 27.4 | 2056 | |
| Pinus radiata LiDAR_2009 (n=82) |
G (m2 ha-1) | LIN (ElCURTmeanCUBE - Intmode - Elp75) | 0.61 | 7.6 | 322 |
| H0 (m) | LIN (Elmax - Elp01 - Elp99 - Ret above 2) | 0.86 | 2.4 | 147 | |
| Wcr (Mg ha-1) | LIN (ElCURTmeanCUBE - Intmode - Elp75 - Intp01) | 0.64 | 7.6 | 1388 | |
| Wst (Mg ha-1) | LIN (Elp99 - ElCURTmeanCUBE) | 0.62 | 34.2 | 1616 | |
| Pinus radiata LiDAR_2011 (n=60) |
G (m2 ha-1) | LIN (ElCURTmeanCUBE - SR) | 0.65 | 5.5 | 158 |
| H0 (m) | EXP (Elmax - SR) | 0.68 | 3.7 | 123 | |
| Wcr (Mg ha-1) | LIN (ElCURTmeanCUBE - SR) | 0.68 | 5.2 | 761 | |
| Wst (Mg ha-1) | EXP (Elmax - SR - SAVI - ElL4) | 0.82 | 17.0 | 869 |
Fig. 2 - Field vs. predicted measured values of basal area (a), dominant height (b), crown biomass (c) and stem biomass (d) for Pinus radiata.
Independent variables related to the height distribution metrics were included in all models (Tab. 3). Intensity metrics appeared as explanatory variables in Eucalyptus spp. (basal area and dominant height), Pinus pinaster (LiDAR_2011 data: basal area, crown biomass and stem biomass) and Pinus radiata (LiDAR_2009 data: basal area and crown biomass - Tab. 3). Landsat derived vegetation indices were explanatory variables in Pinus radiata (LiDAR_2011 data), with Simple Ratio (SR) in basal area, dominant height and crown biomass models, and Soil-Adjusted Vegetation Index (SAVI) in the stem biomass model. The inclusion of Landsat resulted in an increase in the R2 values (from 0.64 to 0.65 for basal area, from 0.64 to 0.68 for dominant height, from 0.66 to 0.68 for crown biomass and from 0.60 to 0.82 for stem biomass), and reductions in RMSE (from 5.6 to 5.5 m2 ha-1 for basal area, from 3.9 to 3.7 m for dominant height, from 5.4 to 5.2 Mg ha-1 for crown biomass and from 25.5 to 17.0 Mg ha-1 for stem biomass) and AIC values (from 161 to 158 for basal area, from 126 to 123 for dominant height, from 767 to 761 for crown biomass and from 899 to 869 for stem biomass).
Tab. 4 displays the model-assisted estimates for each species of the mean values of basal area and dominant height, crown and stem biomass which were obtained by applying the models achieved to the polygons dominated by each analyzed species in the whole study area. The standard errors for estimates of the means were small, ranging from 0.5 to 0.8 m2 ha-1 for basal area, from 0.3 to 0.7 m for dominant height, from 0.6 to 0.8 Mg ha-1 for crown biomass and from 2.5 to 5.2 Mg ha-1 for stem biomass. Bias estimates for the model assisted estimator were also small, from -0.08 to 0.08 m2 ha-1 for basal area, from -0.01 to 1.40 m for dominant height, from -0.01 to 0.01 Mg ha-1 for crown biomass and from -0.01 to 0.03 Mg ha-1 for stem biomass.
Tab. 4 - Estimates mean values (± standard error) of basal area (G) and dominant height (H0), crown biomass (Wcr) and stem biomass (Wst) for each species for the whole study area.
| Main species | G (m2 ha-1) |
H0 (m) |
Wcr (Mg ha-1) |
Wst (Mg ha-1) |
|---|---|---|---|---|
| Eucalyptus spp. | 12.8 ± 0.8 | 19.0 ± 0.7 | 11.4 ± 0.8 | 32.6 ± 5.2 |
| Pinus pinaster | 13.9 ± 0.7 | 14.3 ± 0.4 | 11.0 ± 0.6 | 29.5 ± 2.7 |
| Pinus radiata | 13.6 ± 0.6 | 14.7 ± 0.3 | 14.5 ± 0.6 | 29.9 ± 2.5 |
Discussion
The findings of this study demonstrate that stand dendrometric and biomass variables can be predicted with reasonable precision by using low density LiDAR variables in combination with Landsat data. The importance of linear and exponential models has previously been reported in studies relating LiDAR metrics (combined or not with spectral data) and stand dendrometric and biomass variables ([14], [15]), even for shrub vegetation ([12]). The R2 values obtained in the present study were of the same order of magnitude or slightly smaller than those previously reported for the same species ([16], [14], [15]), probably as a consequence of the use of lower density LiDAR data, or (in the present study), the use of field data from the National Forest Inventory rather than specifically measured field data.
Previous studies have found that height metrics are closely correlated with stand dendrometric and biomass variables for the same species ([14], [15]) and others ([25]). However, these studies were carried out with higher density LiDAR data and specifically measured field data. The most frequent height metric variables observed in our models are maximum height, the cubic mean of the height and diverse percentiles of height distribution. This is consistent with previously reported correlations between stand dendrometric and/or biomass variables and maximum height ([26]) and also between the former and height percentiles ([14], [15], [25]). The most appropriate height metrics reported in the literature widely differ as a likely consequence of differences in the vegetation structure and data processing procedures used ([10]). The close correlation between aboveground biomass values and maximum height and the higher LiDAR percentiles observed in our study may be a result of the regular structure of the forest plantations surveyed ([15]).
Although intensity metrics may not always selected as explanatory variables ([15]), some studies on pine stands have also reported that the inclusion of intensity variables may improve the predictive power of the models, or may even be decisive if used in combination with density or height LiDAR values ([16], [14]).
In this study the least quality of fit observed for Eucalyptus spp. compared to Pinus species is consistent with previous findings ([11]) and is explained by the sparser canopies of Eucalyptus trees, which result in less accurate digital canopy models.
Although there were not previous studies combining LiDAR and satellite data for these species in NW Spain, an improvement in model accuracy by combining both types of data has been previously observed in other tree species ([9], [11]). Some authors only used optical imagery for the first step in vegetation classification in order to account for the dependence of stand dendrometric and biomass estimation on vegetation types ([9]). In the present study, this classification was implemented by using the Spanish Forest Map based on the analysis of aerial photographs. We used Landsat derived vegetation indices as explanatory variables in combination with LiDAR variables to improve the stand dendrometric and biomass models ([11]). In the present study Landsat derived vegetation indices contributed to significantly improve the quality of fit of the model to the data for Pinus radiata (LiDAR_2011 data). The spectral data alone had a small explanatory power, as previously observed for other tree species ([12], [25]). However, the improvement in model performance for Pinus radiata (LiDAR_2011 data) by the inclusion of Landsat derived vegetation indices is consistent with previous studies in which forest stand structure metrics were predicted using a combination of LiDAR and remote sensing imagery ([11], [25]), although the improvement was relatively small in our case.
The standard error of the model-assisted regression estimates were smaller than those obtained by field sampling (simple random sampling estimates - Tab. 1), confirming that remotely sense data made substantial contributions with greater precision, as compared with estimates based only on plot observations ([33]). Mean values of the model-assisted regression values were smaller than those of the plot observations (Tab. 1), likely because field plots were characterized by tree cover > 20% and the presence of trees with dbh > 7.5 cm, while the area covered by remotely sensed data also included locations with smaller tree cover and/or without the presence of trees with dbh > 7.5 cm.
Conclusions
Our findings confirmed the potential of the combined use of freely available remote sensing and regional forest inventories to establish relationships that allow to determine the spatial distribution of both stand dendrometric and aboveground biomass variables of stands dominated by different species in a large area (60 × 60 km) in Galicia. This is the first study combining this freely available information for these species in this area. This information is critical for calibrating and validating biogeochemical models, quantifying carbon fluxes and supporting the United Nations Framework Convention on Climate Change program ([10]).
The future periodicity and availability of this information will enable spatial estimation of stand, biomass and carbon temporal evolution in different vegetation types. The findings of this study support the use of this approach to reduce the cost of forest inventories, thus providing a useful tool enabling stakeholders to map forest stand variables and biomass stocks.
Acknowledgements
This research was funded by Projects of the Conselleria de Medio Rural e do Mar Xunta de Galicia (FEADER 2013/38) and INIA RTA 2014-00011-C06 (FEDER). This work was also co-financed by the INIA and European Social Fund (via a grant awarded to E. Jiménez). We also thank the anonymous reviewer for the useful comments and suggestions, which helped to improve an earlier version of the manuscript.
References
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
CrossRef | Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
CrossRef | Gscholar
Gscholar
Gscholar
Authors’ Info
Authors’ Affiliation
José A Vega
José M Fernández-Alonso
Centro de Investigación Forestal - Lourizán, PO Box 127, 36080 Pontevedra (Spain)
Pablito M López-Serrano
Carlos A López-Sánchez
Facultad de Ciencias Forestales - Universidad Juárez del Estado de Durango (México) Río Papaloapan, Valle del Sur, 34120 Durango, Dgo. (México)
Departmento de Ingeniería de Recursos Naturales y Medio Ambiente, Universidad de Vigo, Campus A Xunqueira, Pontevedra, 36005 (Spain)
Corresponding author
Paper Info
Citation
Jiménez E, Vega JA, Fernández-Alonso JM, Vega-Nieva D, Ortiz L, López-Serrano PM, López-Sánchez CA (2017). Estimation of aboveground forest biomass in Galicia (NW Spain) by the combined use of LiDAR, LANDSAT ETM+ and National Forest Inventory data. iForest 10: 590-596. - doi: 10.3832/ifor1989-010
Academic Editor
Piermaria Corona
Paper history
Received: Jan 20, 2016
Accepted: Mar 12, 2017
First online: May 15, 2017
Publication Date: Jun 30, 2017
Publication Time: 2.13 months
Copyright Information
© SISEF - The Italian Society of Silviculture and Forest Ecology 2017
Open Access
This article is distributed under the terms of the Creative Commons Attribution-Non Commercial 4.0 International (https://creativecommons.org/licenses/by-nc/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Web Metrics
Breakdown by View Type
Article Usage
Total Article Views: 55949
(from publication date up to now)
Breakdown by View Type
HTML Page Views: 45496
Abstract Page Views: 4940
PDF Downloads: 4251
Citation/Reference Downloads: 57
XML Downloads: 1205
Web Metrics
Days since publication: 3294
Overall contacts: 55949
Avg. contacts per week: 118.90
Article Citations
Article citations are based on data periodically collected from the Clarivate Web of Science web site
(last update: Mar 2025)
Total number of cites (since 2017): 14
Average cites per year: 1.56
Publication Metrics
by Dimensions ©
Articles citing this article
List of the papers citing this article based on CrossRef Cited-by.
Related Contents
iForest Similar Articles
Review Papers
Accuracy of determining specific parameters of the urban forest using remote sensing
vol. 12, pp. 498-510 (online: 02 December 2019)
Review Papers
Remote sensing-supported vegetation parameters for regional climate models: a brief review
vol. 3, pp. 98-101 (online: 15 July 2010)
Research Articles
High resolution biomass mapping in tropical forests with LiDAR-derived Digital Models: Poás Volcano National Park (Costa Rica)
vol. 10, pp. 259-266 (online: 23 February 2017)
Review Papers
Analysis of full-waveform LiDAR data for forestry applications: a review of investigations and methods
vol. 4, pp. 100-106 (online: 01 June 2011)
Research Articles
Afforestation monitoring through automatic analysis of 36-years Landsat Best Available Composites
vol. 15, pp. 220-228 (online: 12 July 2022)
Research Articles
Mapping the vegetation and spatial dynamics of Sinharaja tropical rain forest incorporating NASA’s GEDI spaceborne LiDAR data and multispectral satellite images
vol. 18, pp. 45-53 (online: 01 April 2025)
Research Articles
Assessing the availability of forest biomass for bioenergy by publicly available satellite imagery
vol. 11, pp. 459-468 (online: 02 July 2018)
Research Articles
Modeling aboveground carbon in flooded forests using synthetic aperture radar data: a case study from a natural reserve in Turkish Thrace
vol. 17, pp. 277-285 (online: 27 September 2024)
Research Articles
Classification of xeric scrub forest species using machine learning and optical and LiDAR drone data capture
vol. 18, pp. 357-365 (online: 07 December 2025)
Research Articles
Above ground biomass estimation from UAV high resolution RGB images and LiDAR data in a pine forest in Southern Italy
vol. 15, pp. 451-457 (online: 03 November 2022)
iForest Database Search
Search By Author
Search By Keyword
Google Scholar Search
Citing Articles
Search By Author
Search By Keywords
PubMed Search
Search By Author
Search By Keyword