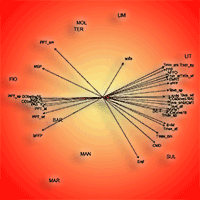
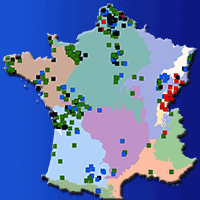
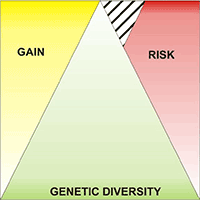
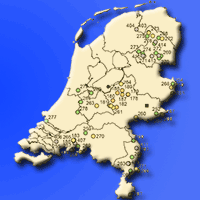
The assessment of seed zones or regions of provenance (RoP) to preserve local adaptation of tree species is an effective tool for the correct management of forest reproductive materials. The RoP for a species or sub-species is the area or group of areas subject to sufficiently uniform ecological conditions in which stands or seed sources show similar phenotypic or genetic characters, taking into account altitudinal boundaries where appropriate. However, the delineation of RoPs is commonly based on estimates of intrinsic environmental homogeneity, mainly climate and/or soil characteristics. The integration of genetic data into RoP maps is an important strategy to obtain a sound tool for managing forest reproductive materials. A study on Quercus suber (cork oak) in Sardinia (Italy) was carried out with the aim of determining ecological regions of provenance, investigating the genetic diversity among populations at the regional scale and identifying possible areas of interest for valorising the available germplasm. Identification of these areas was performed by Reserve Selection Analysis, which allows to identify priority areas by assessing the minimum number of sites required to include all the genetic diversity estimated by genetic analysis. Four spatial clusters were obtained based on environmental data: the northern and northern-eastern parts of the island were included in the Northern RoP; the second RoP covered the western part; and the third RoP enclosed the south-eastern region. The last group was distributed on the central part of the island (Central RoP) and includes the higher elevations. The sampled populations showed a low differentiation among populations and low diversity. According to the Reserve Selection Analysis, four conservation priority areas were identified. These indications can be useful at the local level because these sites can be proposed as stands for seed collection for future plantations.
Keywords
, , , ,
Citation
De Dato G, Teani A, Mattioni C, Marchi M, Monteverdi MC, Ducci F (2018). Delineation of seed collection zones based on environmental and genetic characteristics for Quercus suber L. in Sardinia, Italy. iForest 11: 651-659. - doi: 10.3832/ifor2572-011
Academic Editor
Piermaria Corona
Paper history
Received: Jul 29, 2017
Accepted: Aug 02, 2018
First online: Oct 04, 2018
Publication Date: Oct 31, 2018
Publication Time: 2.10 months
© SISEF - The Italian Society of Silviculture and Forest Ecology 2018
Open Access
This article is distributed under the terms of the Creative Commons Attribution-Non Commercial 4.0 International (https://creativecommons.org/licenses/by-nc/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.

Breakdown by View Type
(Waiting for server response...)
Article Usage
Total Article Views: 49409
(from publication date up to now)
Breakdown by View Type
HTML Page Views: 40221
Abstract Page Views: 4218
PDF Downloads: 3853
Citation/Reference Downloads: 3
XML Downloads: 1114
Web Metrics
Days since publication: 2787
Overall contacts: 49409
Avg. contacts per week: 124.10
Article Citations
Article citations are based on data periodically collected from the Clarivate Web of Science web site
(last update: Mar 2025)
Total number of cites (since 2018): 9
Average cites per year: 1.13
Publication Metrics
by Dimensions ©
Articles citing this article
List of the papers citing this article based on CrossRef Cited-by.
(1)
Aranda I, Castro L, Pardos M, Gil L, Pardos JA (2005)Effects of the interaction between drought and shade on water relations, gas exchange and morphological traits in cork oak (
Quercus suber L.) seedlings. Forest Ecology and Management 210: 117-129.
CrossRef |
Gscholar
(2)
Beatty GE, Brown JA, Cassidy EM, Finlay CMV, McKendrick L, Montgomery WI, Reid N, Tosh DG, Provan J (2015)Lack of genetic structure and evidence for long-distance dispersal in ash (
Fraxinus excelsior) populations under threat from an emergent fungal pathogen: implications for restorative planting. Tree Genetics and Genomes 11 (3): 148.
CrossRef |
Gscholar
(3)
BFW (2017)Austrian regions of provenance. Web site.
Online |
Gscholar
(4)
Boshier D, Broadhurst L, Cornelius J, Gallo L, Koskela J, Loo J, Petrokofsky G, St Clair B (2015)Is local best? Examining the evidence for local adaptation in trees and its scale. Environmental Evidence 4: 311.
CrossRef |
Gscholar
(5)
Bullitta S, Dettori S, Manchinu M, Filigheddu MR, Piluzza G (2011)Characterization of Sardinian cork oak (
Quercus suber L.) genetic resources for economically important traits. Genetic Resources and Crop Evolution 58: 1007-1020.
CrossRef |
Gscholar
(6)
Canu S, Rosati L, Fiori M, Motroni A, Filigheddu R, Farris E (2015)Bioclimate map of Sardinia (Italy). Journal of Maps 11: 711-718.
CrossRef |
Gscholar
(7)
Chapuis M-P, Estoup A (2007)Microsatellite null alleles and estimation of population differentiation. Molecular Biology and Evolution 24: 621-631.
CrossRef |
Gscholar
(8)
Coelho AC, Lima MB, Neves D, Cravador A (2006)Genetic diversity of two evergreen oaks (
Quercus suber L. and
Quercus ilex subsp.
rotundifolia Lam.) in Portugal using AFLP markers. Silvae Genetica 55: 105-118.
CrossRef |
Gscholar
(9)
Corona P, Dettori S, Filigheddu MR, Maetzke F, Scotti R (2005)Site quality evaluation by classification tree: an application to cork quality in Sardinia. European Journal of Forest Research 124: 37-46.
CrossRef |
Gscholar
(10)
Correia B, Valledor L, Meijón M, Rodriguez JL, Dias MC, Santos C, Cañal MJ, Rodriguez R, Pinto G (2013)Is the interplay between epigenetic markers related to the acclimation of cork oak plants to high temperatures? PLoS One 8: e53543.
CrossRef |
Gscholar
(11)
Costa A, Pereira H, Madeira M (2010)Analysis of spatial patterns of oak decline in cork oak woodlands in Mediterranean conditions. Annals of Forest Science 67: 204.
CrossRef |
Gscholar
(12)
Costantini E, Barbetti R, Fantappiè M, Abate G, Lorenzetti R, Napoli R, Marchetti A, Rivieccio R (2014)The soil map of Italy: a hierarchy of geodatabases, from soil regions to sub-systems. In: “GlobalSoilMap: Basis of the Global Spatial Soil Information System” (Arrouays D, McKenzie N, Hempel J, Richer de Forges A, McBratney AB eds). CRC Press Taylor and Francis Group, London, UK, pp. 109-112.
Online |
Gscholar
(13)
Da Costa JSPN (2011)Differentiation and genetic variability in cork oak populations (
Quercus suber L.). PhD thesis, Faculdade de Ciencias Departamento de Biologia Animal, University of Lisbon, Lisbon, Portugal, pp. 120.
Online |
Gscholar
(14)
Daly C, Halbleib M, Smith JI, Gibson WP, Doggett MK, Taylor GH, Curtis J, Pasteris PP (2008)Physiographically sensitive mapping of climatological temperature and precipitation across the conterminous United States. International Journal of Climatology 28: 2031-2064.
CrossRef |
Gscholar
(15)
De Kort H, Mergeay J, Vander Mijnsbrugge K, Decocq G, Maccherini S, Kehlet Bruun HH, Honnay O, Vandepitte K (2014)An evaluation of seed zone delineation using phenotypic and population genomic data on black alder. Journal of Applied Ecology 51: 1218-1227.
CrossRef |
Gscholar
(16)
Demeke T, Sasikumar B, Hucl P, Chibbar RN (1997)Random Amplified Polymorphic DNA (RAPD) in cereal improvement. Maydica 42: 133-142.
Gscholar
(17)
Dow BD, Ashley MV, Howe HF (1995)Characterisation of highly variable (GA/CT)n microsatellites in the bur oak,
Quercus macrocarpa. Theoretical and Applied Genetics 91: 137-141.
CrossRef |
Gscholar
(18)
Ducci F (2015)Genetic resources and forestry in the Mediterranean region in relation to global change. Annals of Silvicultural Research 39: 70-93.
CrossRef |
Gscholar
(19)
Earl DA, Von Holdt BM (2012)STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation of Genetic Resources 4: 359-361.
CrossRef |
Gscholar
(20)
Evanno G, Regnaut S, Goudet J (2005)Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology 14: 2611-2620.
CrossRef |
Gscholar
(21)
Excoffier L, Lischer HEL (2010)Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources 10: 564-567.
CrossRef |
Gscholar
(22)
Ferrara C, Marchi M, Fares S, Salvati L (2017)Sampling strategies for high quality time-series of climatic variables in forest resource assessment. iForest - Biogeosciences and Forestry 10: 739-745.
CrossRef |
Gscholar
(23)
Giorgi F, Lionello P (2008)Climate change projections for the Mediterranean region. Global and Planetary Change 63: 90-104.
CrossRef |
Gscholar
(24)
GRASS Development Team (2017)Geographic Resources Analysis Support System (GRASS) Software, version 7.2. Open Source Geospatial Foundation, Web site.
Online |
Gscholar
(25)
Hammer O, Harper DAT, Ryan PD (2001)PAST: Paleontological statistics software package for education and data analysis. Palaeontologia Electronica 4: 1-9.
Gscholar
(26)
Harris I, Jones PD, Osborn TJ, Lister DH (2014)Updated high-resolution grids of monthly climatic observations - the CRU TS3.10 Dataset. International Journal of Climatology 34: 623-642.
CrossRef |
Gscholar
(27)
Hastie T, Tibshirani R, Friedman J (2001)14 Unsupervised learning. In: “The Elements of Statistical Learning: Data Mining, Inference, and Prediction” (Hastie T, Tibshirani R, Friedman J eds). Springer Open Ltd, New York, USA, pp. 272-280.
CrossRef |
Gscholar
(28)
Hijmans RJ, Guarino L, Cruz M, Rojas E (2001)Computer tools for spatial analysis of plant genetic resources data: 1. DIVA-GIS. Plant Genetic Resources Newsletter 127: 15-19.
Online |
Gscholar
(29)
Hoban S, Schlarbaum S (2014)Optimal sampling of seeds from plant populations for
ex-situ conservation of genetic biodiversity, considering realistic population structure. Biological Conservation 177: 90-99.
CrossRef |
Gscholar
(30)
Hornero J, Gallego FJ, Martinez I, Toribio M (2001)Testing the conservation of
Quercus spp. microsatellites in the cork oak,
Q. suber L. Silvae Genetica 50: 162-167.
Online |
Gscholar
(31)
Hufford KM, Veneklaas EJ, Lambers H, Krauss SL (2016)Genetic delineation of local provenance defines seed collection zones along a climate gradient. AoB PLANTS 8: plv149.
CrossRef |
Gscholar
(32)
INFC (2005)Secondo Inventario Nazionale delle Foreste e dei Serbatoi di Carbonio [2
nd National Inventory of Forests and Carbon Sinks]. Web site. [in Italian]
Online |
Gscholar
(33)
Jakobsson M, Rosenberg NA (2007)CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23: 1801-1806.
CrossRef |
Gscholar
(34)
Jansson G, Danusevicius D, Grotehusmann H, Kowalczyk J, Skrppa T, Krajmerova D, Wolf H (2013)Norway Spruce (
Picea abies (L.) H. Karst.). In: “Forest Tree Breeding in Europe. Current State-of-the-Art and Perspectives” (Paques LE ed). Springer Netherlands, pp. 123-176.
CrossRef |
Gscholar
(35)
Jones TA (2013)When local isn’t best. Evolutionary Applications 6: 1109-1118.
CrossRef |
Gscholar
(36)
Kalinowski ST (2005)HP-RARE 1.0: a computer program for performing rarefaction on measures of allelic richness. Molecular Ecology Notes 5: 187-189.
CrossRef |
Gscholar
(37)
Kampfer S, Lexer C, Glössl J, Steinkellner H (1998)Characterization of (GA)n microsatellite loci from
Quercus robur. Hereditas 129: 183-186.
CrossRef |
Gscholar
(38)
Karp A, Edwards K, Bruford M, Vosman B, Morgante M, Seberg O, Kremer A, Boursot P, Arctander P, Tautz D, Hewitt G (1997)Molecular technologies for biodiversity evaluation: opportunities and challenges. Nature Biotechnology 15: 625-628.
CrossRef |
Gscholar
(39)
Kleinschmit JRG, Kownatzki D, Gregorius H-R (2004)Adaptational characteristics of autochthonous populations consequences for provenance delineation. Forest Ecology and Management 197: 213-224.
CrossRef |
Gscholar
(40)
Konnert M, Fady B, Gömöry D, O’Hara S, Wolter F, Ducci F, Koskela J, Bozzano M, Maaten T, Kowalczyk J (2015)Use and transfer of forest reproductive material in Europe in the context of climate change. European Forest Genetic Resources Programme (EUFORGEN), Bioversity International, Rome, Italy, pp. 75.
Online |
Gscholar
(41)
Kremer A, Ronce O, Robledo-Arnuncio JJ, Guillaume F, Bohrer G, Nathan R, Bridle JR, Gomulkiewicz R, Klein EK, Ritland K, Kuparinen A, Gerber S, Schueler S (2012)Long-distance gene flow and adaptation of forest trees to rapid climate change. Ecology Letters 15: 378-392.
CrossRef |
Gscholar
(42)
Löpez-Aljorna A, Bueno MA, Aguinagalde I, Martín JP (2007)Fingerprinting and genetic variability in cork oak (
Quercus suber L.) elite trees using ISSR and SSR markers. Annals of Forestry Science 64: 773-779.
CrossRef |
Gscholar
(43)
Magalhães AP, Verde N, Reis F, Martins I, Costa D, Lino-Neto T, Castro PH, Tavares RM, Azevedo H (2015)RNA-Seq and gene network analysis uncover activation of an ABA-dependent signalosome during the cork oak root response to drought. Frontiers in Plant Science 6: 1195.
CrossRef |
Gscholar
(44)
Magri D, Fineschi S, Bellarosa R, Buonamici A, Sebastiani F, Schirone B, Simeone MC, Vendramin GG (2007)The distribution of
Quercus suber chloroplast haplotypes matches the palaeogeographical history of the western Mediterranean. Molecular Ecology 16: 5259-5266.
CrossRef |
Gscholar
(45)
Maliouchenko O, Palmé AE, Buonamici A, Vendramin GG, Lascoux M (2007)Comparative phylogeography and population structure of European
Betula species, with particular focus on
B. pendula and
B. pubescens. Journal of Biogeography 34: 1601-1610.
CrossRef |
Gscholar
(46)
Marchi M, Castaldi C, Merlini P, Nocentini S, Ducci F (2015)Stand structure and influence of climate on growth trends of a Marginal forest population of
Pinus nigra spp.
nigra. Annals of Silvicultural Research 39: 100-110.
CrossRef |
Gscholar
(47)
Marchi M, Chiavetta U, Castaldi C, Di Silvestro D, Contu F, Ducci F (2016)Regions of provenance for reproductive materials of the three main forest species of Abruzzi. Journal of Maps 12: 94-97.
CrossRef |
Gscholar
(48)
Modesto IS, Miguel C, Pina-Martins F, Glushkova M, Veloso M, Paulo OS, Batista D (2014)Identifying signatures of natural selection in cork oak (
Quercus suber L.) genes through SNP analysis. Tree Genetics and Genomes 10: 1645-1660.
CrossRef |
Gscholar
(49)
Nanson A (2001)The new OECD scheme for the certification of forest reproductive materials. Silvae Genetica 50: 181-186.
Online |
Gscholar
(50)
Nei M (1973)Analysis of gene diversity in subdivided populations. Proceedings of the National Academy of Sciences USA 70: 3321-3323.
CrossRef |
Gscholar
(51)
Paetkau D, Waits LP, Clarkson PL, Craighead L, Vyse E, Ward R, Strobeck C (1998)Variation in genetic diversity across the range of North American Brown Bears. Conservation Biology 12: 418-429.
CrossRef |
Gscholar
(52)
Peakall R, Smouse PE (2006)GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Molecular Ecology 6: 288-295.
CrossRef |
Gscholar
(53)
Peakall R, Smouse PE (2012)GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28: 2537-2539.
CrossRef |
Gscholar
(54)
Piotti A, Leonardi S, Buiteveld J, Geburek T, Gerber S, Kramer K, Vettori C, Vendramin GG (2012)Comparison of pollen gene flow among four European beech (Fagus sylvatica L.) populations characterized by different management regimes. Heredity 108: 322-331.
CrossRef |
Gscholar
(55)
Pritchard JK, Stephens M, Donnelly P (2000)Inference of population structure using multilocus genotype data. Genetics 155: 945-959.
Online |
Gscholar
(56)
Ramírez-Valiente JA, Lorenzo Z, Soto A, Valladares F, Gil L, Aranda I (2009)Elucidating the role of genetic drift and natural selection in cork oak differentiation regarding drought tolerance. Molecular Ecology 18: 3803-3815.
CrossRef |
Gscholar
(57)
Rebelo AG, Siegfried WR (1992)Where should nature reserves be located in the Cape floristic region, South Africa? Models for the spatial configuration of a reserve network aimed at maximizing the protection of floral diversity. Conservation Biology 6: 243-252.
CrossRef |
Gscholar
(58)
Rosenberg NA (2004)DISTRUCT: a program for the graphical display of population structure. Molecular Ecology Notes 4: 137-138.
CrossRef |
Gscholar
(59)
Rzigui T, Cherif J, Zorrig W, Khaldi A, Nasr Z (2017)Adjustment of photosynthetic carbon assimilation to higher growth irradiance in three-year-old seedlings of two Tunisian provenances of Cork Oak (
Quercus suber L.). iForest - Biogeosciences and Forestry 10: 618-624.
CrossRef |
Gscholar
(60)
Sáenz-Romero C, Lamy JB, Ducousso A, Musch B, Ehrenmann F, Delzon S, Cavers S, Chalupka W, Dagdas S, Hansen JK, Lee SJ, Liesebach M, Rau HM, Psomas A, Schneck V, Steiner W, Zimmermann NE, Kremer A (2017)Adaptive and plastic responses of
Quercus petraea populations to climate across Europe. Global Change Biology 23: 2831-2847.
CrossRef |
Gscholar
(61)
Sedda L, Delogu G, Dettori S (2011)Forty-four years of land use changes in a Sardinian cork oak agro-silvopastoral system: a qualitative analysis. The Open Forest Science Journal 4: 57-66.
CrossRef |
Gscholar
(62)
Soto A, Lorenzo Z, Gil L (2003)Nuclear microsatellite markers for the identification of
Quercus ilex L. and
Q. suber L. hybrids. Silvae Genetica 52: 63-66.
Online |
Gscholar
(63)
Steinkellner H, Fluch S, Turetschek E, Lexer C, Streiff R, Kremer A, Burg K, Glössl J (1997)Identification and characterization of (GA/CT)n - microsatellite loci from
Quercus petraea. Plant Molecular Biology 33: 1093-1096.
CrossRef |
Gscholar
(64)
Toumi L, Lumaret R (1998)Allozyme variation in cork oak (
Quercus suber L.): the role of phylogeography and genetic introgression by other Mediterranean oak species and human activities. Theoretical and Applied Genetics 97: 647-656.
CrossRef |
Gscholar
(65)
UKFC (2017)Regions of provenance and native seed zones. Forestry Commission, UK, Web site.
Online |
Gscholar
(66)
Van Zonneveld M, Scheldeman X, Escribano P, Viruel MA, Van Damme P, Garcia W, Tapia C, Romero J, Sigueñas M, Hormaza J (2012)Mapping genetic diversity of cherimoya (
Annona cherimola Mill.): application of spatial analysis for conservation and use of plant genetic resources. PLoS One 7: e29845.
CrossRef |
Gscholar
(67)
Vander Mijnsbrugge K, Bischoff A, Smith B (2010)A question of origin: where and how to collect seed for ecological restoration. Basic and Applied Ecology 11: 300-311.
CrossRef |
Gscholar
(68)
Varela MC (1997)Regions of provenance for Cork oak in Portugal. In: “
Quercus suber Network. Report of the 3
rd and 4
th meetings” (Turok J, Varela MC, Hansen C eds). Sassari (Italy) 9-12 June 1996 and Almoraima (Spain) 20-22 Feb 1997. International Plant Genetic Resources Institute, Rome, Italy, pp. 37-42.
Gscholar
(69)
Vinceti B, Loo J, Gaisberger H, Van Zonneveld MJ, Schueler S, Konrad H, Kadu CA, Geburek T (2013)Conservation priorities for
Prunus africana defined with the aid of spatial analysis of genetic data and climatic variables. PLoS One 8: e59987.
CrossRef |
Gscholar
(70)
Wang T, Hamann A, Spittlehouse DL, Murdock TQ (2012)ClimateWNA-high-resolution spatial climate data for western North America. Journal of Applied Meteorology and Climatology 51: 16-29.
CrossRef |
Gscholar
(71)
Whitlock R, Hipperson H, Thompson DBA, Butlin RK, Burke T (2016)Consequences of
in-situ strategies for the conservation of plant genetic diversity. Biological Conservation 203: 134-142.
CrossRef |
Gscholar
(72)
Williams MI, Dumroese RK (2013)Preparing for climate change: forestry and assisted migration. Journal of Forestry 111: 287-297.
CrossRef |
Gscholar
(73)
Zucca GM (2011)Molecular and phenoptypic characterization of
Quercus suber L. and
Pinus uncinata R. populations in the Mediterranean basin. PhD thesis, Scienze dei Sistemi Agrari e Forestali e delle Produzioni Alimentari, University of Sassari, Sassari, Italy, pp. 124.
Gscholar