Clonal structure and dynamics of peripheral Populus tremula L. populations
iForest - Biogeosciences and Forestry, Volume 7, Issue 3, Pages 140-149 (2014)
doi: https://doi.org/10.3832/ifor0971-007
Published: Jan 13, 2014 - Copyright © 2014 SISEF
Research Articles
Abstract
Native Populus tremula L. populations at the south-western limit of the species’ range are threatened by the anthropic reduction of their habitats and by climate change. In these areas, P. tremula forms small and multiclonal populations. Knowledge of the structures and dynamics of these populations is essential to design effective conservation programs. The clonal spatial and ramet age distributions of three natural populations in north-western Spain were studied. Trees with diameter > 3 cm at breast height were mapped and their age identified. Multilocus genotypes based on 11 isoenzymatic loci were obtained for a sample of 90-106 evenly distributed trees per population. Clonal assignment of the remaining trees in each stand was interpolated using geostatistical methods. All three stands were multiclonal, with higher clonal diversity than that reported in northern populations, and no genotype was detected in more than one stand. Most clones exhibited aggregated distributions, but some scattered clones were also found. The clonal structures of the stands are consistent with the hypothesis that recruitment is absent inside the stand. A combined analysis of ages and genotypes revealed cycles of regrowth in which each clone experienced varying degrees of success in generating new ramets. These degrees of success could be the consequence of varying abilities for vegetative propagation among genotypes. Geostatistical interpolation can be a useful tool for the elaboration of clonal maps, thus reducing the costs of sampling and genotyping, but further research is needed to define the optimum scale of sampling for accurate clonal assignment. For P. tremula conservation programs based on clonal material, small plantations with relatively few clones (approximately 10) provide an appropriate strategy if the metapopulation is maintained at a sufficient level of diversity. The set of clones used for plantations and ex-situ collections should consider both the most abundant and the least frequent clones to avoid a biased selection.
Keywords
Populus tremula, Clonal Structure, Genetic Diversity, Population Dynamics, Conservation Plans
Introduction
Native Populus forests have suffered a dramatic reduction in their natural distribution area, and are now considered one of the most threatened ecosystems in Europe (see [28] and [51] for P. nigra, [11] and [43] for P. tremula). These species occur in a wide variety of habitats, support very high levels of biodiversity ([15]) and have high structural complexity ([7]). Their populations experience frequent disturbances (mostly periodic floods, fires and strong winds) that create opportunities for regeneration ([25]). In recent decades, poplar forests are threatened by human activities that modify population dynamics by restricting the disturbance regimes or destroying the natural habitat, resulting in a strong reduction of its distribution areas.
Spanish populations of Populus tremula L. (European aspen) represent the southwestern limit of the species’ global distribution. In the context of climate change, southern extremes of P. tremula range are highly sensitive areas where important environmental changes are expected to occur. For example, in Spain, increases in temperature, drought and forest fires are expected ([30], [38]). In addition to climate change, browsing by herbivores ([39]), insect incursion and root diseases have been associated with aspen decline ([34]). In Spain, induced forest fires and changes in land use are also probable causes for the rapid disappearance of many populations. Consequently, the European aspen is listed as a priority species in the Spanish Strategy for the Conservation and Sustainable Use of Forest Genetic Resources ([40]). Knowledge of the genetic structure and dynamics of natural populations is central for detecting changes and developing appropriate management and conservation strategies. In particular, southern peripheral populations are likely the most sensitive to the effects of global change ([36]).
P. tremula is a pioneer dioecious species. In Spain, it usually grows in small groups. Flowering typically occurs every year, in contrast to areas with cool and wet summers such as Scotland, where flowering is rare ([11]). The germination potential of seeds is high ([54]), although seeds are short-lived and require special conditions for germination and establishment. Life expectancy of P. tremula trees is relatively short (less than 100 years); however, this species has the great ability to produce root suckers, allowing clones to persist for long periods ([2], [37]). Temperatures above 25 °C and the availability of light appear to stimulate sucker production ([24]).
The vegetative propagation ability of P. tremula and the small sizes of its populations suggest that its populations are monoclonal. However, in a study of five populations in northern Spain using isozyme markers, López-de-Heredia et al. ([32]) observed a multiclonal structure in each population. Escudero & Sierra-de-Grado ([13]) observed both multiclonal and monoclonal compositions in 44 populations in the Douro basin. In addition, Suvanto & Latva-Karjanmaa ([49]) identified a multiclonal composition in a set of stands in an old-growth managed forest in Finland. A multiclonal composition presumably involves genotypes originating from different seeds. DeWoody et al. ([9]) demonstrated that the majority of intrastand variation in Populus tremuloides Michx. was likely due to the presence of different clones rather than somatic mutation in old trees. However, once a population is founded by the first set of seeds, the balance between new recruitment and root suckers remains unclear, as does the interplay among different clones within the stand.
Two different growth forms or strategies have been distinguished in clonal plants ([33]): spreading (guerrilla form) and clumping ramets (phalanx form). Growth forms condition the distribution and colonisation of the site in patches with varying extents of aggregation. The size of a clone can be affected by both its vegetative propagation ability (in terms of the number of ramets produced) and its clonal growth strategy ([47]). According to Widen et al. ([53]), the most vigorous or more advantageously placed clones develop roots spreading in a linear mode as long superficial roots in the early stages, whereas clones in more advanced stages expand in globoid forms by branching their roots.
Populus spp. seeds disperse and germinate in areas cleared by recent disturbances, establish new stands, produce new travelling seeds and, after some time, are extinguished by disturbances such as strong winds or flooding. The above dynamic determines the existence of metapopulations, which are considered genetically interconnected collections of isolated individuals, clumps and stands that colonise and become extinct locally while maintaining a long-term presence over a wider area ([29], [49], [9]). In particular, P. tremula forms metapopulations over a wide range of environmental conditions; it can be found along riverbanks and colonising eroded rocky slopes. The age structure of the ramets can offer valuable data for understanding the clone dynamics and the evolution of the stand ([20], [2], [37], [31]). If one stand expands from the germination of different seeds and the subsequent vegetative propagation of each individual, several clones can be expected in young populations. At the first stage, with space readily available, the number of ramets per clone would increase depending on the regrowth capacity of each clone. After this initial expansion, certain clones may decrease in frequency or disappear due to competition, whereas those better adapted or having a higher ability for vegetative propagation will expand. Thus, fewer clones with an unbalanced distribution are expected in old populations compared with a higher clonal diversity in young populations. A combined analysis of the spatial distribution of clones and of the ages of the clone ramets provides an approach to charting the historical development of the stand based on living trees.
Geostatistical methods are based on the assumption that data are spatially correlated. Geostatistics applied to multi-allele or multilocus genotypes have been demonstrated to be useful for the analysis of fine-scale genetic structures in natural populations ([50], [48], [10]). In clonal populations, we can assume that the proximity and relative positions of the trees indicate the probability of a tree belonging to a given clone. Experimental evidence for this assumption is presented by Suvanto & Latva-Karjanmaa ([49]) and Mock et al. ([37]). This spatial correlation allows data interpolation and calculation of the probability that a tree belongs to one of multiple different clones based on their spatial coordinates. This spatial prediction ability allows a probabilistic approach to the analysis of clonal distribution in the stand, which is a useful tool for the optimization of sampling design and cost reduction of sampling and genotyping in conservation and management programs.
The aim of this work was to analyse the genetic diversity, spatial distribution of the clones and ages of the ramets in three populations of P. tremula in the south-western limit of its range. A better understanding of the dynamics of this type of population can help to optimise conservation and management strategies.
Materials and methods
Populations
Three populations in the north of Palencia province (north-western Spain) were selected (Tab. 1) as being representative of the different developmental stages of P. tremula in terms of stem age. Monte Aguilar is a stand of young, high-density thin trees, the majority of which have grown since a forest fire 26 years before current survey. Monte Aguilar is surrounded by Quercus pyrenaica Willd. and Quercus petraea (Mattuschka) Liebl. and, in some areas, is bounded by dense heathlands of Erica spp. The Congosto population is a mature stand with a low crown ratio within a Q. pyrenaica forest near a stream bed. There is historical evidence of a larger population of P.tremula in this area ([35]), now reduced to a small stand of old trees. The Tremaya population shows a stage of intermediate development between the other two populations. Tremaya population grows on a steep slope surrounded by forest composed of Fagus sylvatica L. and Q. petraea. The stands are not in close proximity to each other: the distance from Tremaya to Congosto and Monte Aguilar is 33 and 17 km, respectively, and it is 37 km from Congosto to Monte Aguilar.
Tab. 1 - Location and characteristics of the studied stands: latitude, longitude, altitude (m a.s.l.), slope, orientation, number of inventoried trees with diameters at breast height > 3 cm, area of the stand (m2), basal area (m2/ha) and expected age of the population.
| Stand | Lat / Long | Altitude (m a.s.l.) |
Slope / Orientation |
Inventoried trees |
Area (m2) |
Basal area (m2/ha) |
Expected age |
|---|---|---|---|---|---|---|---|
| Monte Aguilar |
42°53’40’’ N 4°14’50’’ W |
1080 | 10-40% S |
1301 | 3000 | 9.38 | young |
| Tremaya | 42°58’40’’ N 4°27’39’’ W |
1170 | 70-100% N |
644 | 11308 | 8.99 | intermediate |
| Congosto | 42°42’20’’ N 4°37’54’’ W |
1040 | 10-30% NE |
269 | 5400 | 18.64 | old |
Inventory and sampling
All the aspen stems with a diameter at breast height (DBH) > 3 cm were mapped in each population. The high stand density made it difficult to use total station or GPS; thus, a vertex hypsometer and a compass were used. The DBH was measured using a digital calliper.
Tree age was determined by dendrochronological analysis in samples of 93, 60 and 51 stems in Monte Aguilar, Tremaya and Congosto, respectively. Trees sampled for age measurements were randomly selected from the whole population. Increment cores were collected at the stem base, and the age of each tree was determined with a magnifying glass by adding a mixture of 0.1 N HCl and 1% fluoroglucine to improve the visibility of the tree rings. Finally, age estimates for remnant stems were based on the best fit non-linear regression model for each population, using the DBH as the explanatory variable.
In total, 93 stems in Monte Aguilar, 90 stems in Tremaya and 106 stems in Congosto were used for genetic analysis. We collected winter twigs to use the dormant buds for isozyme extraction. Trees were selected evenly throughout the stand using a sampling mesh with a variable side length depending on the stand size. In the two larger stands (Tremaya and Congosto), sampling was performed in two stages. In the first step, we collected approximately one third of the samples (30 and 29 evenly distributed samples in Tremaya and Congosto, respectively). Subsequent to genetic analysis, a second set of samples was collected in the following winter, intensifying the sampling of: (a) zones of the stands where several clones had been detected (to better establish the boundaries between the clones); (b) zones where trees were in different age classes compared with the previous sample; and (c) the area surrounding trees belonging the less abundant clones. Several trees were resampled in the second phase to test the consistency of the markers used.
Genetic analysis
Horizontal starch gel electrophoresis was used to assay six isoenzymatic systems (LAP, GOT, ADH, IDH, SKDH, 6PGD). Starch gels (12%) were prepared, and sucrose was added (3.5%) to facilitate migration. In total, three to four buds from each tree were crushed in several drops of extraction buffer to extract enzymes. The staining procedures and buffers followed those described by Alba & Agúndez ([1]) and Wendel & Weeden ([52]). An extensive review of literature on Populus species was used to interpret the results for different enzyme systems ([16], [42], [8], [1], [32]).
The software package Popgene v.1.31 ([55]) was used to estimate diversity parameters based on the allelic frequencies (number of alleles per locus - Na, effective number of alleles per locus - Ne), percentage of polymorphic loci and expected and observed heterozygosity in each population ([41]).
We performed a multilocus analysis, assuming that trees sharing the same genotype belong to the same clone, to assess the number of genetically different trees and to map the clones. We also assumed that each different genotype originated from a different seed; the probability that these assumptions are true depends on the number of loci and variability of isoenzymatic systems used in clonal identification. DeWoody et al. ([9]) demonstrated that the ability of isoenzyme markers to dinstinguish clones in P. tremuloides is similar to that using DNA markers as microsatellites, provided an appropriate number of markers. The discriminating power of the isoenzyme analysis was calculated by the Aspinwall & Christian ([4]) estimator. This estimator provides the mean probability of the samples that share the same genotype for a given number of loci belonging to the same clone (eqn. 1):
where Q is a tree from a certain population, N is the number of trees in the population, M is the number of polymorphic loci in the population, XDQ is the number of trees with the same isozyme pattern at locus D as tree Q, and PD is the total number of trees analysed for isozyme locus D.
We calculated the genetic diversity (G/N, where G is the number of different multilocus genotypes and N is the number of samples) and the number of clones per unit area (G/A, where A is the area in hectares). In addition, we calculated an estimate of the genotypic evenness E ([14], [3]) independent of the sample size.
Geostatistical analysis
After proper assignment of genotyped trees to the various clones, a geostatistical analysis was carried out ([18], [6]) to estimate the probability of clone membership for the non-genotyped trees. This method results in a map of the stand’s clonal distribution, based on the most probable clonal assignments. Therefore, we conducted a structural analysis by calculating experimental variograms independently for each clone. For these experimental variograms, the semivariance of a pair of trees separated by a distance h equals zero if the two trees belong to the same clone; otherwise, the semivariance equals 0.5. Experimental variograms of this type provide a similar interpretation as classical indicator variograms ([18]).
In variograms, the semivariance increases up to a certain distance that is called the range. In our study, the range defines the separation distance of trees within which the spatial autocorrelation is lost (equivalent to the average spatial size of the clone). Theoretically, the semivariance should decrease to zero at zero distance (the origin of the variogram graph). However, in practice there is often a nonzero semivariance at the origin known as the nugget effect, which can be caused by spatial variability at distances below the minimum sampling interval and/or measurement errors (see also [6]).
The probability of clone membership for non-genotyped trees was computed using ordinary kriging based on a previously adjusted variogram model. We adjusted the kriging neighbourhood based on the variogram range of each separate clone. A non-genotyped ramet was assigned to the most likely clone using the kriging probabilities of clone membership. All geostatistical analyses were performed in the Isatis environment ([17]).
Results
Genetic analysis
A total of 11 loci were interpreted, 9 of which were polymorphic, with 24 alleles. Isozyme analysis allowed to identify a total of 30 multilocus genotypes in 289 genotyped stems sampled from all populations (Tab. 2). This number of distinct genotypes is relatively low compared with the number of trees, consistent with the strategy of clonal species. None of the three analysed stands were found to be monoclonal (8 clones in Monte Aguilar, 13 clones in Tremaya and 9 clones in Congosto). The three most abundant clones in each stand accounted for 63%, 65% and 91% of the genotyped stems from Monte Aguilar, Tremaya and Congosto, respectively. None of the genotypes were detected in more than one stand.
Tab. 2 - Number of heterozygous loci and relative frequencies of the multilocus genotypes identified in the studied populations.
| Stand | Genotype | Frequency (%) |
Heterozygous Loci |
|---|---|---|---|
| Monte Aguilar | 27 | 37 | 1 |
| 28 | 14 | 2 | |
| 25 | 12 | 2 | |
| 26 | 10 | 3 | |
| 24 | 8 | 3 | |
| 23 | 8 | 2 | |
| 29 | 8 | 1 | |
| 30 | 3 | 2 | |
| Tremaya | 1 | 24 | 3 |
| 4 | 23 | 3 | |
| 3 | 18 | 2 | |
| 2 | 11 | 2 | |
| 11 | 7 | 0 | |
| 9 | 6 | 1 | |
| 10 | 3 | 1 | |
| 5 | 2 | 2 | |
| 6 | 2 | 2 | |
| 7 | 1 | 2 | |
| 8 | 1 | 1 | |
| 12 | 1 | 1 | |
| 13 | 1 | 1 | |
| Congosto | 16 | 39 | 3 |
| 15 | 27 | 2 | |
| 14 | 25 | 3 | |
| 21 | 3 | 3 | |
| 17 | 2 | 1 | |
| 18 | 1 | 2 | |
| 19 | 1 | 2 | |
| 20 | 1 | 2 | |
| 22 | 1 | 4 |
In Monte Aguilar, 8 genotypes and 15 alleles were observed. For Congosto and Tremaya, the use of a two-stage sampling processes resulted in an increase more than 3-fold in the number of sampled trees per population. As a result, the number of genotypes found in Tremaya increased from 7 in the first step to 13 in the second step and from 6 to 9 in Congosto. The total number of alleles identified increased from 16 to 21 in Tremaya and 17 to 18 in Congosto. The discriminating power of the isozyme analysis ranged from 0.793 for Congosto to 0.893 for Monte Aguilar.
Monte Aguilar contained the fewest distinct genotypes (G=8) but had frequencies that were more balanced and had the highest number of genotypes per ha (G/A=26.7 vs. 11.8 and 18.0 in Tremaya and Congosto, respectively). Tremaya contained the greatest genetic and genotypic diversity (Tab. 3). The population of Congosto had the lowest values for the three parameters used to estimate genotypic diversity, and the population was dominated by a limited number of clones (ramets of the three most abundant clones represented 91% of the tree population) (Tab. 2).
Tab. 3 - Diversity parameters for each population. (N): number of sampled trees; (Na): mean number of alleles per locus; (Ne): effective number of alleles per locus; (P): % of polymorphic loci; observed (Ho) and expected (He) Nei’s heterozygosity (He); (G): total number of multilocus genotypes; (G/N): relative number of multilocus genotypes; (E): modified Simpson’s index; (G/A): number of genotypes per ha.
| Stand | N | Na | Ne | P | Ho | He | G | G/N | E | G/A |
|---|---|---|---|---|---|---|---|---|---|---|
| Monte Aguilar | 93 | 1.444 | 1.225 | 44.44 | 0.191 | 0.146 | 8 | 0.086 | 0.89 | 26.7 |
| Tremaya | 90 | 2 | 1.514 | 88.89 | 0.354 | 0.27 | 13 | 0.144 | 0.873 | 11.8 |
| Congosto | 106 | 1.778 | 1.566 | 66.67 | 0.297 | 0.256 | 9 | 0.085 | 0.751 | 18 |
Age and clonal structure
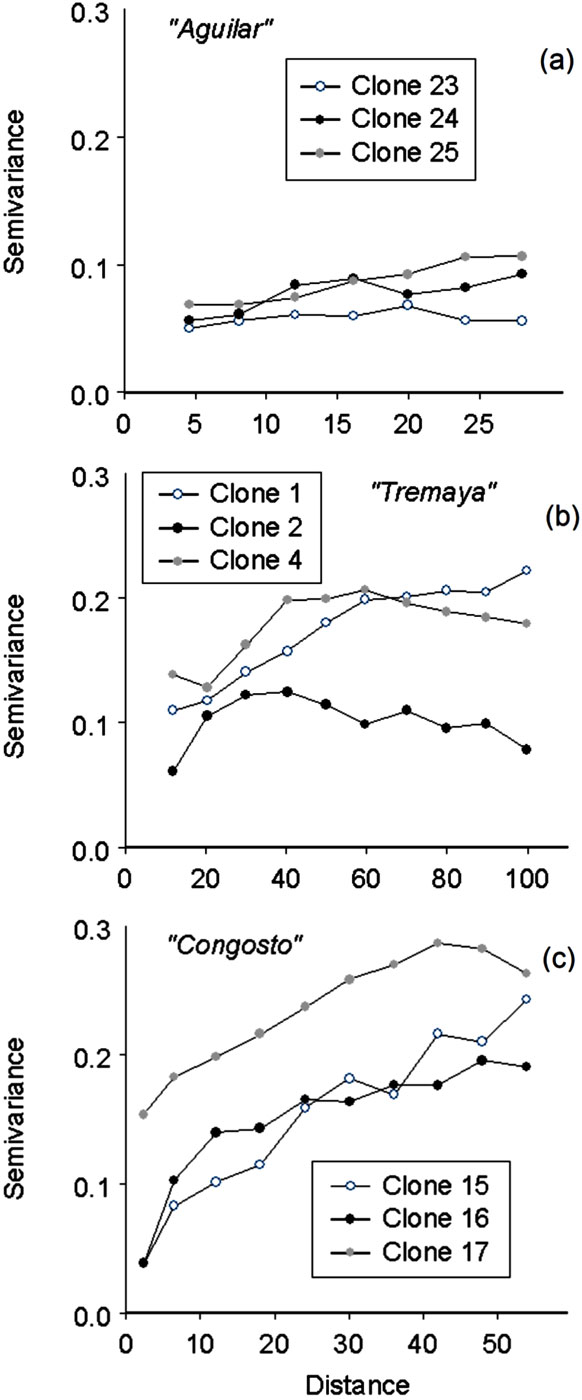
Spatial size of clones (as estimated by the variogram range) varied from 28 to 100 m and from 48 to 80 m at Tremaya and Congosto, respectively, whereas smaller variogram ranges (approximately 15 to 35 m) were observed at Monte Aguilar. The nugget effect variances for eight clones ranged from 0% (clones 15 and 16 - Fig. 1) to 100% (variograms not shown). A pure nugget effect within a variogram suggests the absence of spatial correlation for the specific clone distribution. Variograms of this type were obtained from the analysis of clones with a reduced number of ramets. Prediction probabilities of clone membership for these clones were realised using pure nugget effect variogram models. The minimum variogram range (clone length) that our geostatistical analysis was able to define was 4 m.
Fig. 1 - Experimental variograms for nine clones (of 30) in the three analysed populations of wild aspen: (a) Monte Aguilar; (b) Tremaya; (c) Congosto.
The average clone sizes expressed as the number of ramets per clone were 12, 7 and 12 for Monte Aguilar, Tremaya and Congosto, respectively, when only the genotyped trees were taken into account. However, if we consider the genotypes estimated by geostatistical analysis, the average clone sizes in Monte Aguilar, Tremaya and Congosto were 30, 42 and 163, respectively. The larger clones that were observed consisted of 476, 78 and 302 (genotyped + estimated) ramets, and the percentage of clones consisting of only one ramet was 0%, 23% and 44% in Monte Aguilar, Tremaya and Congosto, respectively.
The following regression models were developed to estimate the age of the trees from which no cores were extracted, in each population:
- Monte Aguilar (r2=0.59): Age = 12.96 + exp (0.1895 · DBH)
- Tremaya (r2=0.6): Age = exp (2.952 + 0.0227 · DBH)
- Congosto (r2=0.87): Age = 6.297 · DBH0.664
From here onwards, results concerning the age of trees in each population refer both to measured plus estimated data, including Fig. 2 to 5, unless otherwise indicated.
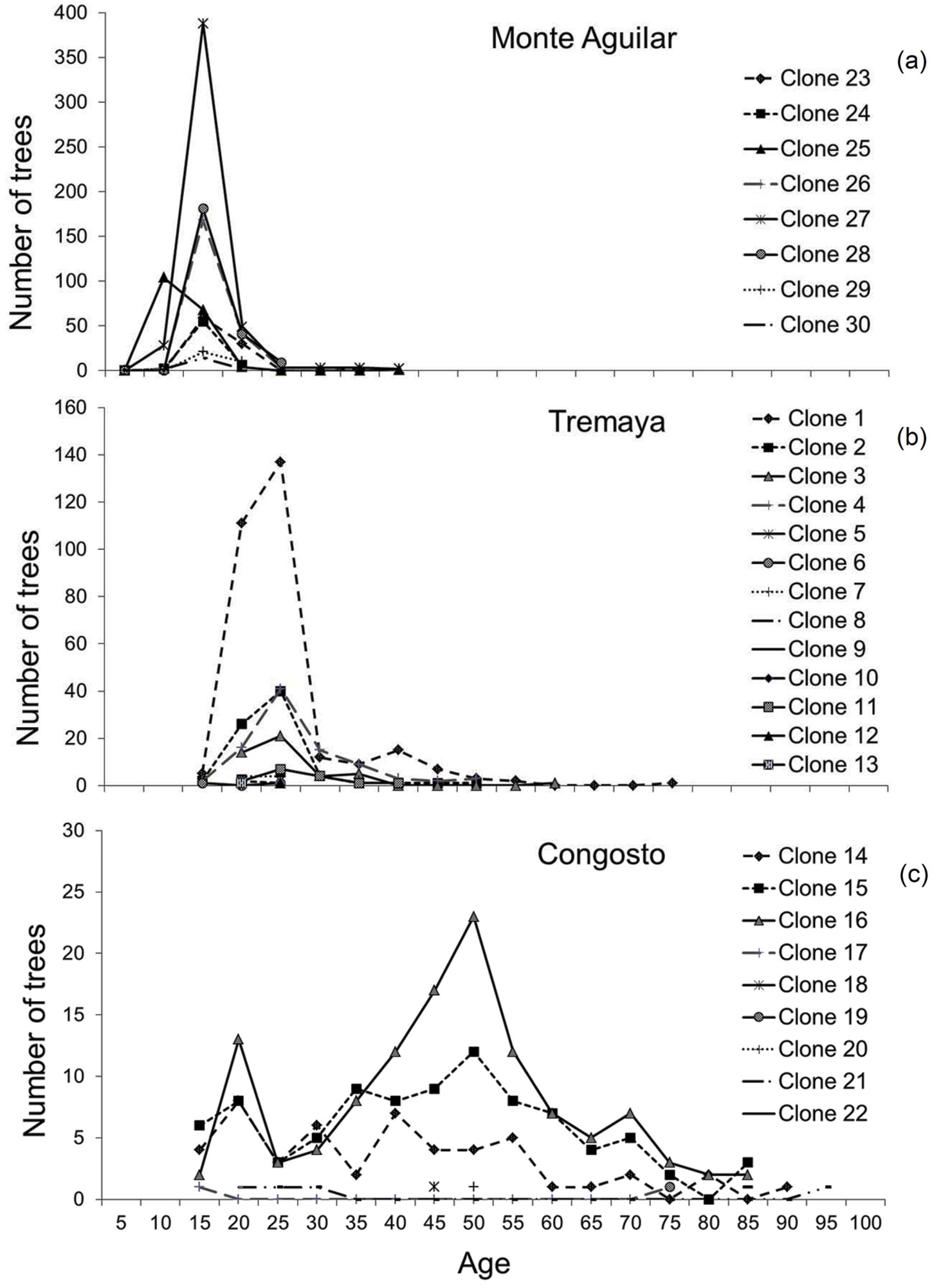
Fig. 2 - Age distributions of the ramets (measured plus estimated data by regression models) for each clone and stand. (a) Monte Aguilar; (b) Tremaya; (c) Congosto.
The ramet age distribution (Fig. 2) shows that Monte Aguilar was an even-aged stand. In contrast, Tremaya showed a dominant age class, although older trees were found within the most widespread clones. In Congosto, the age class distribution was more irregular. The expansion of clones occurs in waves of sprouting, causing certain age classes to be more abundant; clones have varying degrees of success in sprouting and generating new ramets in each wave of regrowth. In Congosto, where the oldest trees were located, ramets formed over a period of 80 years were observed in several clones.
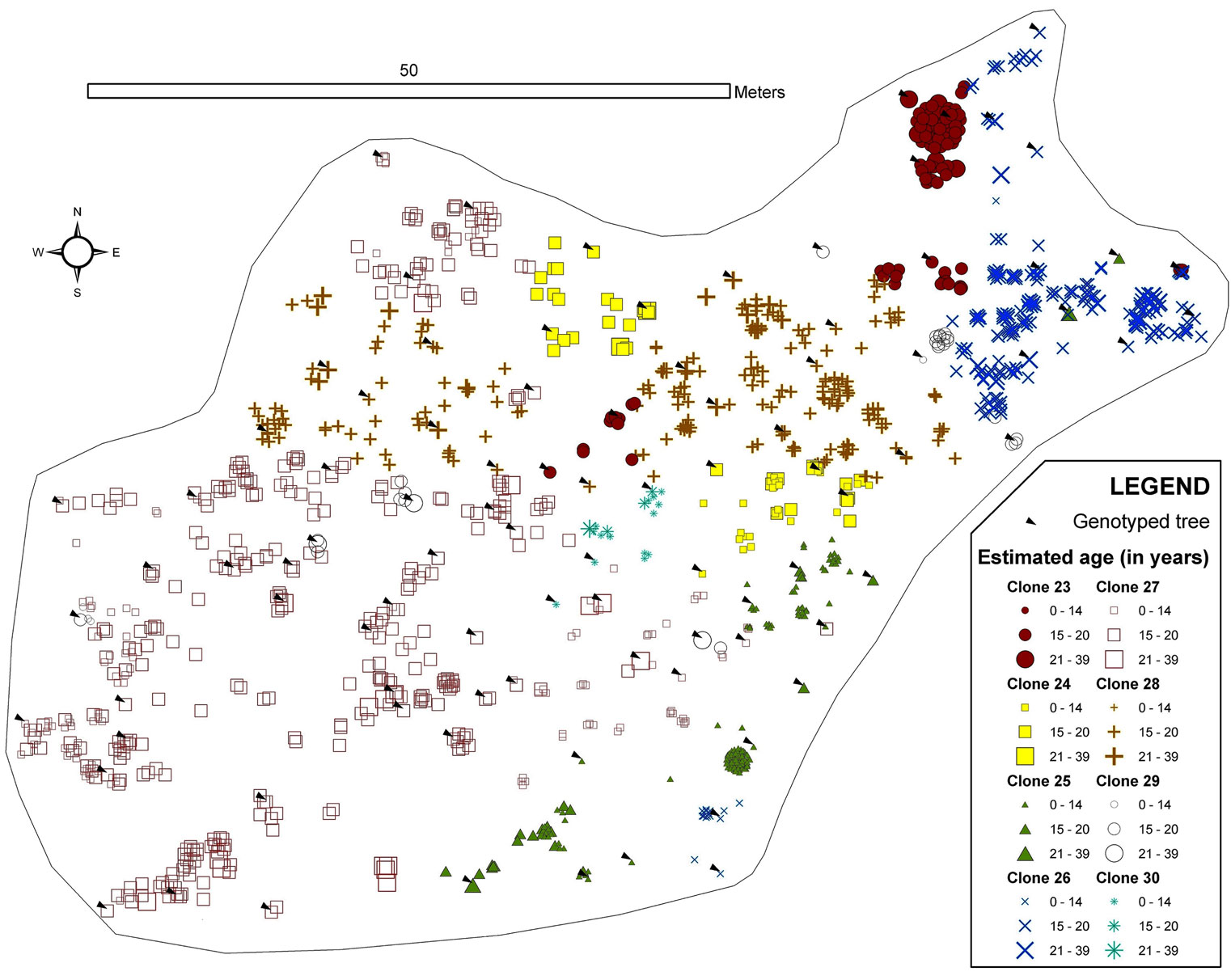
Monte Aguilar was found to contain trees greater than 30 years old (located to the north and east of the stand). Specifically, the dendrochronological analysis revealed two trees that were 36 and 38 years old. A phalanx type of clonal strategy (i.e., ramets that are close together or clustered) was displayed at Monte Aguilar, although there was a slight interdigitation (Fig. 3). In this stand, the predominant age for clone ramets was 16 years.
Fig. 3 - Map of the spatial distribution of clones in the Monte Aguilar stand. All trees with DBH > 3cm were inventoried and georeferenced. The symbols and colours indicate the clonal assignments of each tree: genotyped trees are indicated by arrows, and the genotypes of the trees without arrows were interpolated by geostatistical methods. The size of each symbol indicates the age class of the tree. The symbols for two or more trees can overlap, and some small trees may be hidden.
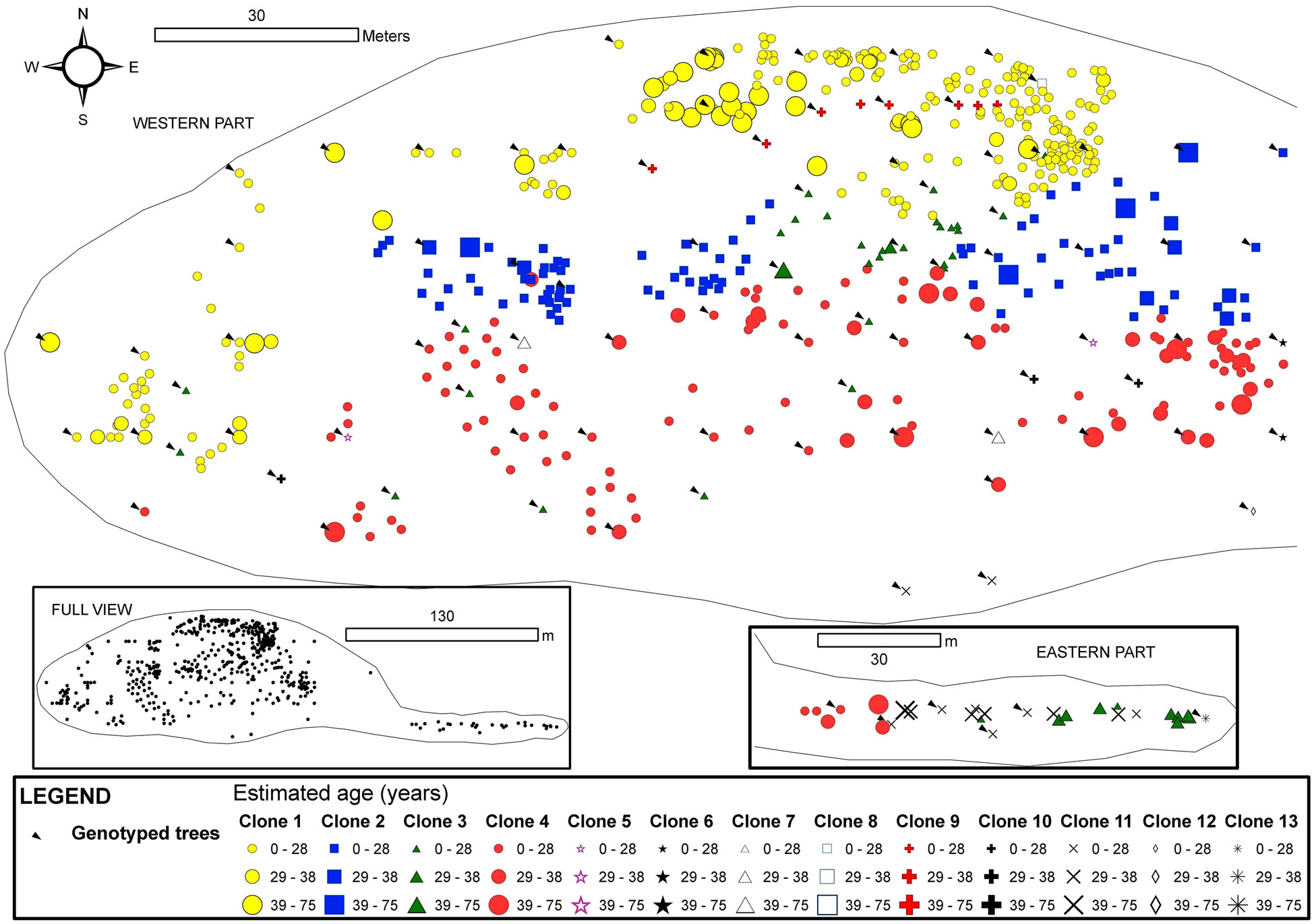
Clones in Tremaya showed a preferential west-east orientation, and the clones were located approximately parallel to the slope contour lines (Fig. 4). In this stand, the average ramet age of the different clones was 28 years, although some trees were up to 70 years old. Tremaya is characterised by a relatively high number of genotypes, with the three most abundant genotypes accounting for 65% of the total number of genotyped trees within the stand. One of the three most abundant clones varied (clone 2 instead of clone 3) based on geostatistical estimations, and these three clones accounted for 86% of the total trees within the stand. Clone 3 was scattered throughout the stand and lacked spatial autocorrelation. Younger trees were found in areas of stand expansion near unvegetated sites to the south-west (clone 1) and west (clone 4).
Fig. 4 - Map of the spatial distribution of clones in the Tremaya stand. All trees with DBH > 3 cm were inventoried and georeferenced. The symbols and colours indicate the clonal assignments of each tree: genotyped trees are indicated by arrows, and the genotypes of the trees without arrows were interpolated by geostatistical methods. The size of each symbol indicates the age class of the tree. The symbols for two or more trees can overlap, and some small trees may be hidden.
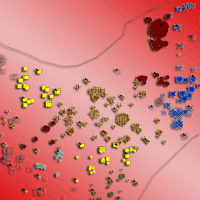
The Congosto stand (Fig. 5) presented a phalanx strategy typical of populations in the last stages of development, where clones appear to form contact groups but do not intermix. However, the borders between clones were delineated by the statistical model. Clone 16 was the most abundant (Tab. 2), located in the central part of the stand and surrounded by clones 14 and 15 with intermediate extension. Several minor clones (18, 19, 20 and 21) were situated in a small area to the south of the stand. An area of greater clonal variability was identified where genotypes 16, 17, 18, 20 and 21 are found. The average age of the clone ramets was 43 years with a mode of 50 years. The oldest tree (nearly 100 years old) was observed at the center of the stand, while the youngest tree was found in the periphery. Asexual propagation in the stand was shown to originate from three clones (14, 15 and 16).
Fig. 5 - Map of the spatial distribution of clones in the Congosto stand. All trees with DBH > 3cm were inventoried and georeferenced. The symbols and colours indicate the clonal assignments of each tree: genotyped trees are indicated by arrows, and the genotypes of the trees without arrows were interpolated by geostatistical methods. The size of each symbol indicates the age class of the tree. The symbols for two or more trees can overlap, and some small trees may be hidden.
Discussion and conclusions
Genetic diversity
The genetic diversity, in terms of the number of clones per ha per population (26.7, 11.8 and 18.0 - Tab. 3), was generally higher than in northern populations, including reports of 4.6, 4.6 and 12.7 clones per ha per population in France ([8]), Scotland ([11]) and Finland ([49]), respectively. The average clone size, expressed as the number of ramets per clone, was higher in our populations (see results) than in aforementioned studies, e.g., 9.8 ([8]), 1.8 ([11]) and 2.1 ([49]) ramets per clone. In this present study, both the number of clones per ha and the number of ramets per clone were higher than in previous studies, indicating that there was a greater density in the Monte Aguilar, Tremaya and Congosto populations. However, data comparison among the above studies is difficult due to the variations in sampling strategies and intensities. The relatively high number of clones per population found for P. tremula contrasts with those of P. alba and P. x canescens in the same river basin (Douro), where only one genotype has been found within an extensive area ([45]).
Peripheral populations are often characterized by low genetic diversity due to isolation, genetic drift, natural selection and inbreeding ([27], [12], [19]). However, the diversity levels found in the studied populations are higher than expected. Furthermore, we have to take into account the fact that the sampling effect and the discriminant power of the set of isozymes (which is high but not 100%) may lead to an underestimation of the actual genetic diversity (i.e., levels of diversity in the studied populations could be even higher than what we found). Better conditions for flowering are found in Spain than in northern countries ([11]), which could favor a higher rate of sexual reproduction and gene flow, resulting in greater seed availability. These results enhance the relevance of the conservation of P. tremula populations in the studied area.
Large environmental differences in the areas covered by each of the three population studied may affect the diversity parameters obtained. In addition, the presence of trees older than the last forest fire (occurred 26 years ago) in Monte Aguilar clearly indicates that the population establishment precedes the fire. It is likely that fire nearly destroyed this population, although part of the root system survived. Certain clones could have disappeared due to this disturbance, although a relatively large number of clones relative to the stand area were preserved. Another consequence of the fire may have been the stimulation of root sucker production, which led to a high tree density (4336 trees/ha) and an abundant representation of the surviving clones (88% of clones with at least eight ramets and no single-ramet clones). In contrast, single-ramet clones represented 31% and 44% of the analysed trees at Tremaya and Congosto, respectively. Previous studies reported single-ramet clone percentages of 16% ([11]), 57% ([8]) and 64% ([49]). The increased number of alleles identified in the second phase of sampling is not proportional to the sampling effort, as suggested by Kimura & Crow ([26]), but indicates a scenario of diminishing returns in which each new unit of investment in the sampling effort returns a lower gain in terms of the number of alleles found ([5]). However, the increase in the number of genotypes was greater than that of alleles, and the increases were lower in the older and smaller population of Congosto than in Tremaya. These results suggest that, although a sample size of 40-50 trees may be sufficient for capturing the allelic diversity of this type of stand, larger samples of 100 or more trees are required for better depicting its clonal composition.
Spatial distribution of clones
In the studied populations, compact clonal patches were common, although some clones appeared to present one dominant direction of expansion, as displayed by clone 15 in Congosto or clones 1, 2 and 4 in Tremaya. However, the scattered distribution of old ramets within the areas occupied by these clones makes it difficult to define a direction of clone spreading and suggests that the origin of the clone (ortet) could have disappeared long ago.
Some clones were scattered throughout the stand and lacked spatial autocorrelation (such as clone 3 in Tremaya or clone 29 in Monte Aguilar). This pattern suggests a clonal growth following the guerrilla strategy instead of the usual phalanx strategy, or such clones could be in recession, i.e., the clone previously exhibited a greater range of expansion but is now being replaced by its neighbors. The high interconnectivity of roots by intra- and interclonal grafting, as found in trembling aspen by Jelínková et al. ([22]), could be an underlying explanation for the existence of separated clumps of ramets, because segments of roots can remain alive for a long time after tree death as they are grafted to the roots of other clones. Alternatively, this hypothetical clone could comprise more than one genotype that was not detected by the set of isoenzymes used in this study. At the western part of the stand, there was a separated row of trees bordering a meadow that contained ramets of clone 3 and others; the origin of the stand may have been anthropogenic, serving to provide shade for animals.
The preferential west-east distribution of clones in Tremaya is possibly due to the topography of the site, which has a steep north-south orientated slope. Root growth (and thus clonal expansion) may be limited by topographical factors, such as slope steepness, sudden changes in slope aspect or the presence of stream beds. Such limitations of expansion can be considered when designing sampling strategies for capturing genetic diversity ([23]).
Combining genotyping of a sample of trees with geostatistical interpolation allowed to obtain a map with the most likely spatial distribution of clones in the stand. This method can result in significant financial savings through the assignment of single stems to clones. However, clones with a single or reduced number of ramets had null or weak spatial correlations, which makes it difficult to estimate the number of ramets through geostatistical analysis. Further research is necessary to define the optimum scale of sampling for an accurate clonal assignment in different scenarios with various frequencies of small clones.
Population dynamics and conservation plans
In the three studied populations, clonal expansion was found to occur in open areas, such as gaps within the stand, or by spreading to adjacent areas. No new genotypes were located in these areas, consistent with a lack of regeneration by seeds within an established stand. In the central zone of the Congosto stand, there was an internal area with young ramets of clone 16, which may have been an area in which a disturbance opened a window of opportunity in the established stand. Importantly, clone ramets, rather than new genotypes, were found within this window, even in the second phase of sampling when the area was resampled with greater intensity. Given the historical references to the existence of this population ([35]), as well as the probable absence of recruitment from inside the stand, it is possible that the present clones originated more than 200 years ago.
In stands where several clones and cohorts of ramets coexist, clones comprised of mature trees only and with little or no regrowth capacity, are more likely to disappear in the event of further disturbances or the death of old trees. In addition, some human actions (such as selective cutting of larger trees) can contribute to the loss of this type of clone.
Management and conservation plans should promote the in situ regeneration of natural populations and their natural dynamics ([29], [44]). In the context of metapopulations, the importance of genetic diversity and the need to preserve genetically interconnected populations gives relevance to small clusters of trees that can contribute to population dynamics by exchanging pollen and seeds ([44]). In European aspen populations at the limit of their range and facing conservation difficulties, the establishment of new stands, though small, may be an effective method of contributing to both diversity and gene flow. To establish new stands in management planning, two approaches are possible: (i) the planting of ramets from several clones; or (ii) the planting of trees from seeds collected from unrelated parents. Although the latter approach introduces greater diversity and allows for the generation of new genotypes that may cope better with future environmental conditions, vegetative propagation is the most frequent approach in tree nurseries of Spain and other European countries ([54], [46]). If vegetative propagation is used, the number of clones must be sufficient to maintain adequate clonal diversity. In addition, an appropriate mixture of male and female clones is required for successful sexual reproduction. This permits high levels of sexual reproduction and sufficient seed production, thus favoring the maintenance of natural processes. Previous studies on Populus nigra L. ([21]) suggest that a minimum of 50 distinct clones is required to avoid genetic risks of inbreeding and a loss of genetic variability, if the population is in genetic contact with other populations. However, 75 distinct clones are required if the likelihood of genetic exchange with other populations is moderate, and 100 distinct clones are needed if the population is completely isolated. However, our results, similar to those of other studies of natural P. tremula stands in Spain ([32], [13]), have shown the presence of a reduced number of clones per stand (between one and 13 per population), and each clone is usually found in only one stand. This result indicates that a limited number of clones is adequate in local plantations if the entire metapopulation has sufficient genetic diversity. This concept is especially interesting in regions where the natural representation of the species is marginal, traditional use for industrial purposes is limited and its introduction in reforestation is as an accessory species only.
The distribution of the number of ramets per clone in each population is sharply unbalanced. Further research is needed to address whether differences in the number of ramets per clone correlate to differences in vegetative propagation ability. Assuming that a large number of ramets implies a robust ability for regrowth and good adaptation, the most abundant clones are interesting candidates for use in the reforestation of degraded areas or sites where rapid revegetation is needed, providing also a good basis for breeding programs. However, because a significant part of the variability of a population relies on less abundant clones, in plantations maintained for conservation purposes or in ex situ collections, it is also important to ensure that less abundant clones are well represented to avoid a biased selection. The key for implementing these recommendations is the availability of sufficient and adequate forest reproductive material. Genetic resource collections of indigenous aspen, established by ex situ conservation measures, can serve as a starting point for the production of reproductive material. The clones that are most abundant in natural stands should be preferentially included in ex situ collections because they are likely to be well adapted and, therefore, would provide a good basis for breeding programs. To provide a robust reforestation plan, the number of ramets per clone should be balanced in newly established plantations. To achieve positive results, two strategies can be adopted: (i) planting ramets of different clones in a random mix that would give an advantage to the more competitive clones; and (ii) planting a mosaic of monoclonal plots that would favor clones with slower expansion rates.
Acknowledgments
We are grateful to three anonymous reviewers for their suggestions on an earlier version of the manuscript. This work was supported by the Consejería de Educación y Cultura of the Junta de Castilla y León (Regional Government), projects VA019A08 and VA 123/02. DC coordinated the genetic analysis; PM analyzed the results and discussion with DC and RS; IV, JC and JV conducted the fieldwork and analysis of Tremaya, Congosto and Monte Aguilar, respectively; NN performed the geostatistical analysis; RS supervised the fieldwork and coordinated the research project.
References
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Gscholar
Authors’ Info
Authors’ Affiliation
Inés Villamediana
Josu Ciriza
Javier Villar
Rosario Sierra-de-Grado
Department of Plant Production and Forest Resources, Universidad de Valladolid, ETSIIAA, Avda. De Madrid 44, 34071 Palencia (Spain)
Pablo Martínez-Zurimendi
Rosario Sierra-de-Grado
Sustainable Forest Management Research Institute UVa-INIA (Spain)
ECOSUR, Villahermosa (México)
Escuela Técnica Superior de Ingenieros de Montes, Universidad Politécnica de Madrid (Spain)
Corresponding author
Paper Info
Citation
Cristóbal D, Martínez-Zurimendi P, Villamediana I, Ciriza J, Villar J, Nanos N, Sierra-de-Grado R (2014). Clonal structure and dynamics of peripheral Populus tremula L. populations. iForest 7: 140-149. - doi: 10.3832/ifor0971-007
Academic Editor
Andrea Piotti
Paper history
Received: Feb 12, 2013
Accepted: Sep 30, 2013
First online: Jan 13, 2014
Publication Date: Jun 02, 2014
Publication Time: 3.50 months
Copyright Information
© SISEF - The Italian Society of Silviculture and Forest Ecology 2014
Open Access
This article is distributed under the terms of the Creative Commons Attribution-Non Commercial 4.0 International (https://creativecommons.org/licenses/by-nc/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
Web Metrics
Breakdown by View Type
Article Usage
Total Article Views: 58257
(from publication date up to now)
Breakdown by View Type
HTML Page Views: 48494
Abstract Page Views: 3490
PDF Downloads: 4630
Citation/Reference Downloads: 35
XML Downloads: 1608
Web Metrics
Days since publication: 4485
Overall contacts: 58257
Avg. contacts per week: 90.93
Article Citations
Article citations are based on data periodically collected from the Clarivate Web of Science web site
(last update: Mar 2025)
Total number of cites (since 2014): 5
Average cites per year: 0.42
Publication Metrics
by Dimensions ©
Articles citing this article
List of the papers citing this article based on CrossRef Cited-by.
Related Contents
iForest Similar Articles
Research Articles
Clonal structure and high genetic diversity at peripheral populations of Sorbus torminalis (L.) Crantz.
vol. 9, pp. 892-900 (online: 29 May 2016)
Technical Reports
Population genetic structure of Platanus orientalis L. in Bulgaria
vol. 4, pp. 186-189 (online: 11 August 2011)
Commentaries & Perspectives
The genetic consequences of habitat fragmentation: the case of forests
vol. 2, pp. 75-76 (online: 10 June 2009)
Technical Reports
Conservation and use of elm genetic resources in France: results and perspectives
vol. 13, pp. 41-47 (online: 03 February 2020)
Research Articles
Patterns of genetic diversity in European beech (Fagus sylvatica L.) at the eastern margins of its distribution range
vol. 10, pp. 916-922 (online: 10 December 2017)
Research Articles
Tree encroachment dynamics in heathlands of north-west Italy: the fire regime hypothesis
vol. 3, pp. 137-143 (online: 27 September 2010)
Research Articles
Genetic diversity of core vs. peripheral Norway spruce native populations at a local scale in Slovenia
vol. 11, pp. 104-110 (online: 31 January 2018)
Review Papers
Genetic diversity and forest reproductive material - from seed source selection to planting
vol. 9, pp. 801-812 (online: 13 June 2016)
Research Articles
Fine-scale spatial genetic structure in a multi-oak-species (Quercus spp.) forest
vol. 8, pp. 324-332 (online: 05 September 2014)
Review Papers
Biodiversity assessment in forests - from genetic diversity to landscape diversity
vol. 2, pp. 1-3 (online: 21 January 2009)
iForest Database Search
Search By Author
Search By Keyword
Google Scholar Search
Citing Articles
Search By Author
Search By Keywords
PubMed Search
Search By Author
Search By Keyword